Simulating fMRI Data
Bradley R. Buchsbaum
2026-04-15
Source:vignettes/a_08_simulation.Rmd
a_08_simulation.RmdIntroduction to fMRI Data Simulation
When developing or validating an fMRI analysis pipeline, you need
data where the ground truth is known – real fMRI data doesn’t come with
known effect sizes or noise parameters. Simulation fills this gap. The
fmrireg package offers several functions to simulate fMRI
data with varying levels of complexity:
-
simulate_bold_signal: Simulates clean BOLD responses for multiple experimental conditions -
simulate_noise_vector: Generates realistic fMRI noise with temporal autocorrelation, drift, and physiological components -
simulate_simple_dataset: Combines signal and noise for a complete dataset based on SNR -
simulate_fmri_matrix: Creates multiple time series with shared event timing but column-specific variation in parameters
This vignette demonstrates how to use these functions to create realistic fMRI simulations for various purposes.
Simulating Clean BOLD Signals
Let’s start with simulate_bold_signal, which generates a
clean hemodynamic response signal for multiple experimental
conditions.
# Simulate 3 conditions with different amplitudes
sim <- simulate_bold_signal(ncond = 3, amps = c(1.0, 1.5, 2.0), TR = 2)
# Extract the data
time <- sim$mat[,1] # First column contains time
signals <- sim$mat[,-1] # Other columns contain condition signalsWe reshape the data into long format so ggplot2 can map each condition to a separate color.
df <- data.frame(
Time = time,
Cond1 = signals[,1],
Cond2 = signals[,2],
Cond3 = signals[,3]
)
df_long <- tidyr::pivot_longer(df, cols = c(Cond1, Cond2, Cond3),
names_to = "Condition",
values_to = "Response")
ggplot(df_long, aes(x = Time, y = Response, color = Condition)) +
geom_line() +
theme_minimal() +
labs(title = "Simulated BOLD Signals for Three Conditions",
x = "Time (seconds)",
y = "BOLD Response",
color = "Condition") +
scale_color_brewer(palette = "Set1")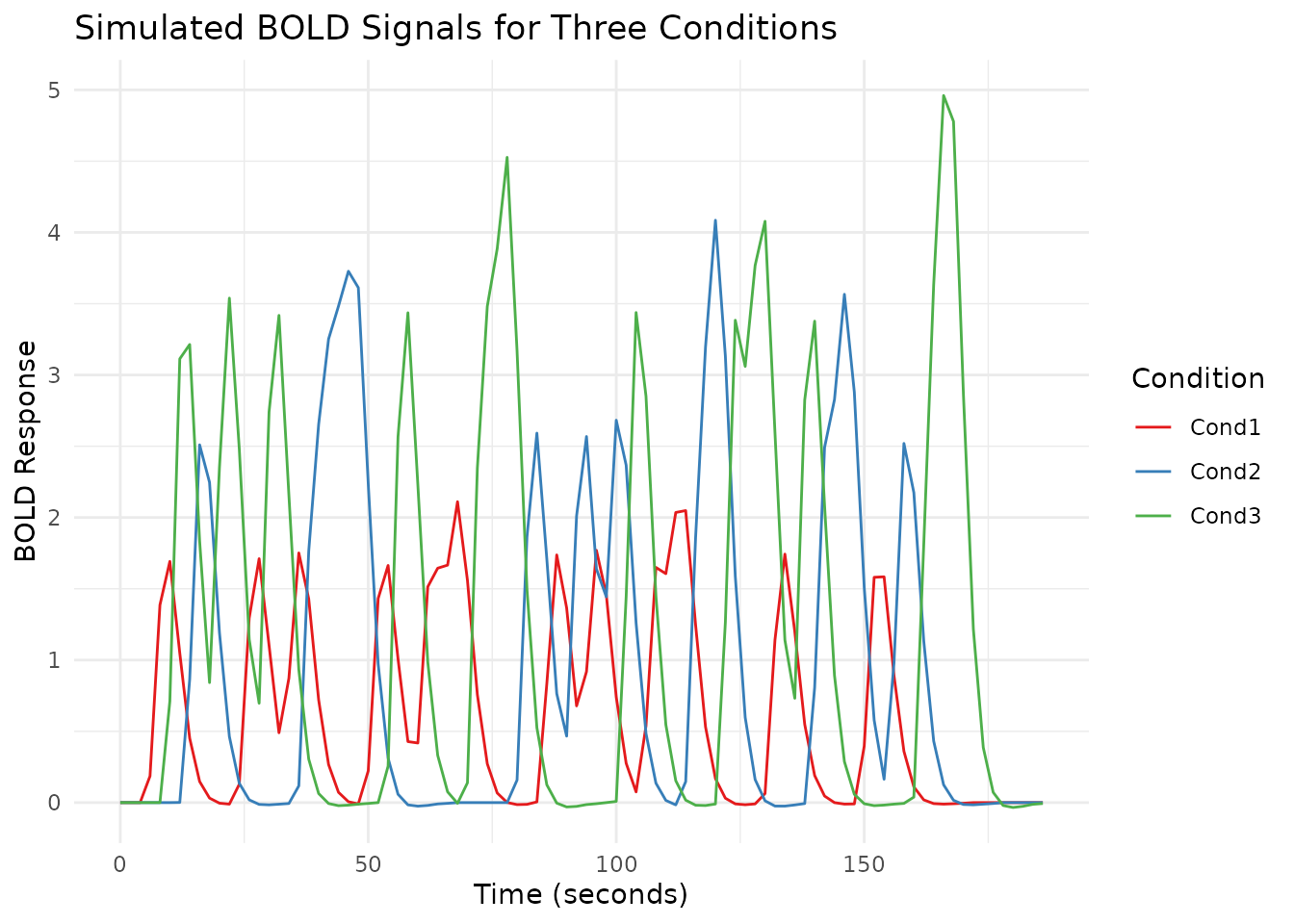
Adding dashed vertical lines at each event onset makes the relationship between stimulus timing and the hemodynamic response easier to see.
ggplot(df_long, aes(x = Time, y = Response, color = Condition)) +
geom_line() +
geom_vline(xintercept = sim$onset, linetype = "dashed", alpha = 0.3) +
theme_minimal() +
labs(title = "Simulated BOLD Signals with Event Onsets",
x = "Time (seconds)",
y = "BOLD Response",
color = "Condition") +
scale_color_brewer(palette = "Set1")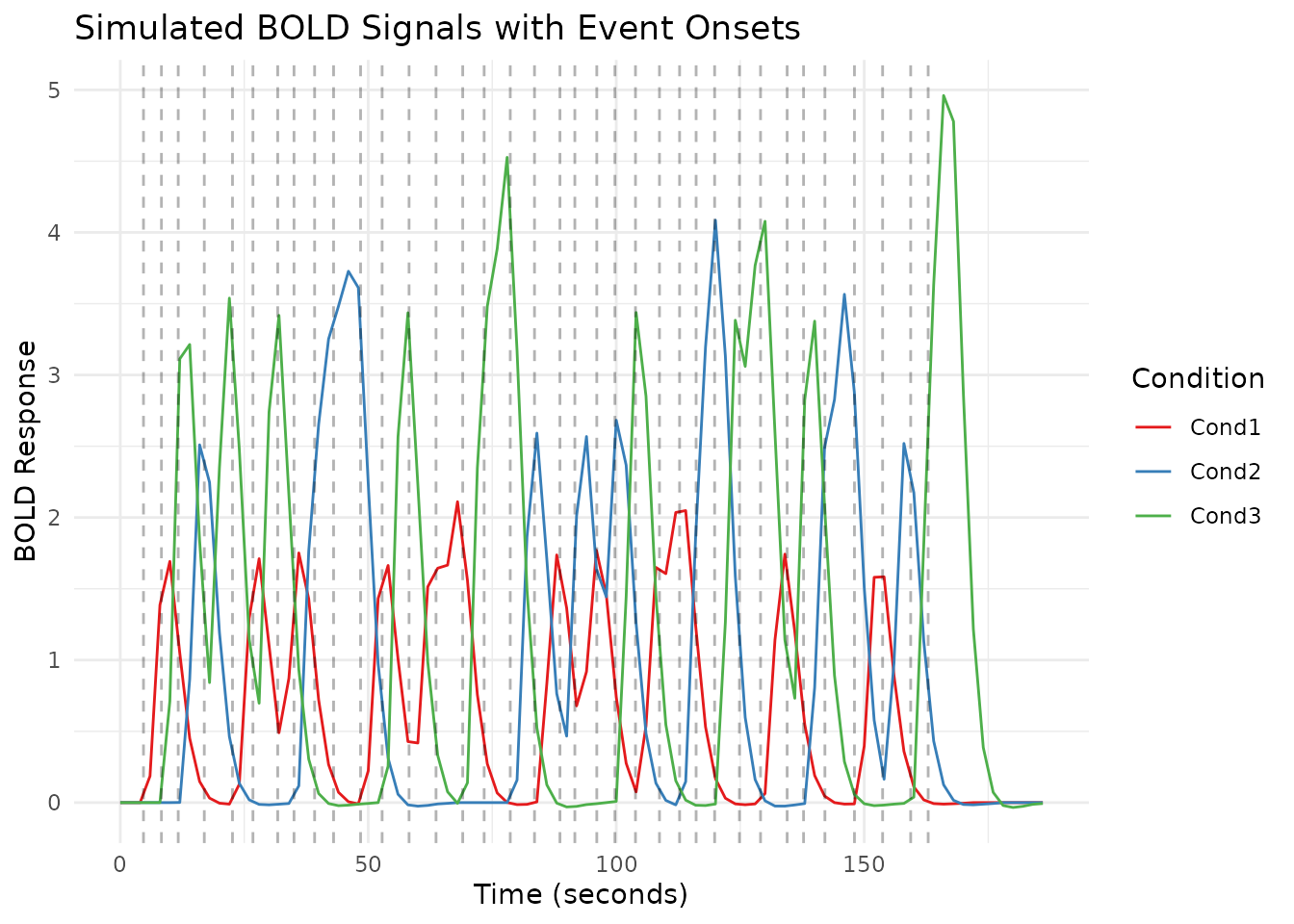
The function returns a list containing: -
onset: Event onset times -
condition: Condition labels for each event
- mat: Matrix with time points and BOLD
responses for each condition
You can control: - Number of conditions (ncond) - Number
of repetitions per condition (nreps) - HRF shape
(hrf) - Amplitudes for each condition (amps) -
Inter-stimulus interval range (isi) - Amplitude variability
(ampsd)
Simulating Realistic fMRI Noise
The simulate_noise_vector function generates realistic
fMRI noise by combining various noise sources common in real fMRI data.
To understand its components, let’s simulate and visualize them
separately and combined.
First, pure white noise – random fluctuations with no temporal structure.
noise_white <- simulate_noise_vector(n_timepoints, TR = TR,
ar = 0, ma = 0,
drift_amplitude = 0, physio = FALSE, sd = 1)Next, ARMA noise introduces temporal autocorrelation, which makes the signal “smoother” than white noise.
noise_arma <- simulate_noise_vector(n_timepoints, TR = TR,
ar = c(0.6), ma = c(0.3),
drift_amplitude = 0, physio = FALSE, sd = 1)Scanner drift is a slow oscillation that shifts the baseline over the course of a run.
drift_freq <- 1/128
drift_amplitude <- 2
noise_drift <- drift_amplitude * sin(2 * pi * drift_freq * time)Physiological noise adds quasi-periodic fluctuations at respiratory and cardiac-like frequencies.
noise_cardiac <- 1.5 * sin(2 * pi * 0.3 * time)
noise_respiratory <- 1.0 * sin(2 * pi * 0.8 * time)
noise_physio <- noise_cardiac + noise_respiratoryFinally, we combine the ARMA, drift, and physiological components to mimic what a real scanner would produce.
noise_combined <- noise_arma + noise_drift + noise_physio
noise_df <- data.frame(
Time = time,
White_Noise = noise_white,
ARMA_Noise = noise_arma,
Drift_Component = noise_drift,
Physiological_Noise = noise_physio,
Combined_Noise = noise_combined
)
noise_long <- tidyr::pivot_longer(noise_df,
cols = -Time,
names_to = "NoiseType",
values_to = "Signal")
noise_long$NoiseType <- factor(noise_long$NoiseType,
levels = c("White_Noise", "ARMA_Noise",
"Drift_Component", "Physiological_Noise",
"Combined_Noise"),
labels = c("White Noise", "ARMA Noise",
"Drift Component", "Physiological Noise",
"Combined Noise"))
ggplot(noise_long, aes(x = Time, y = Signal, color = NoiseType)) +
geom_line() +
facet_wrap(~NoiseType, ncol = 1, scales = "free_y") +
theme_minimal() +
theme(legend.position = "none") +
labs(title = "Simulated fMRI Noise Components: Time Courses",
x = "Time (seconds)",
y = "Signal Amplitude")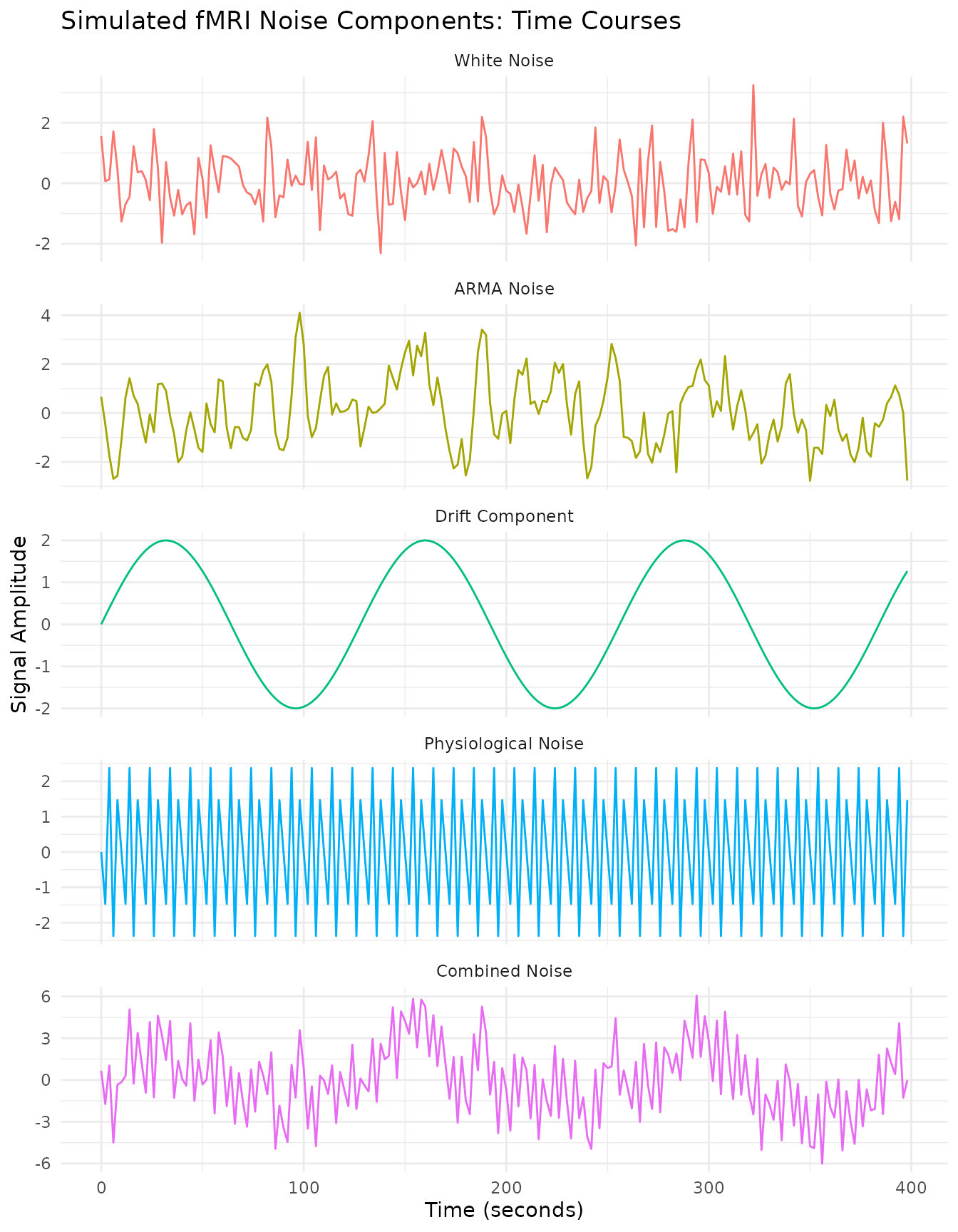
Power Spectrum Analysis
Power spectra reveal the frequency content of each noise type. White noise has a flat spectrum, while ARMA noise concentrates power at lower frequencies.
spec_white <- calculate_spectrum(noise_white, TR)
spec_arma <- calculate_spectrum(noise_arma, TR)
spec_drift <- calculate_spectrum(noise_drift, TR)
spec_physio <- calculate_spectrum(noise_physio, TR)
spec_combined <- calculate_spectrum(noise_combined, TR)
spec_white$NoiseType <- "White Noise"
spec_arma$NoiseType <- "ARMA Noise"
spec_drift$NoiseType <- "Drift Component"
spec_physio$NoiseType <- "Physiological Noise"
spec_combined$NoiseType <- "Combined Noise"
spec_df <- rbind(spec_white, spec_arma, spec_drift, spec_physio, spec_combined)
spec_df$NoiseType <- factor(spec_df$NoiseType,
levels = c("White Noise", "ARMA Noise",
"Drift Component", "Physiological Noise",
"Combined Noise"))We also compute a high-resolution drift spectrum from a longer signal to better resolve the low-frequency peak.
n_long <- 1024
time_long <- seq(0, (n_long - 1) * TR, by = TR)
drift_long <- drift_amplitude * sin(2 * pi * drift_freq * time_long)
spec_drift_long <- calculate_spectrum(drift_long, TR)
spec_drift_long$NoiseType <- "Drift Component (High Resolution)"Each panel below shows the power spectrum of a different noise component. The dashed green line in the drift panel shows the high-resolution version.
ggplot() +
geom_line(data = spec_df, aes(x = Frequency, y = Power, color = NoiseType)) +
geom_line(data = subset(spec_drift_long, Frequency <= 0.05),
aes(x = Frequency, y = Power), color = "darkgreen", linetype = "dashed") +
theme_minimal() +
facet_wrap(~NoiseType, ncol = 1, scales = "free_y") +
theme(legend.position = "none") +
labs(title = "Power Spectra of Different Noise Components",
x = "Frequency (Hz)",
y = "Power") +
scale_color_brewer(palette = "Set1") +
coord_cartesian(xlim = c(0, 0.25))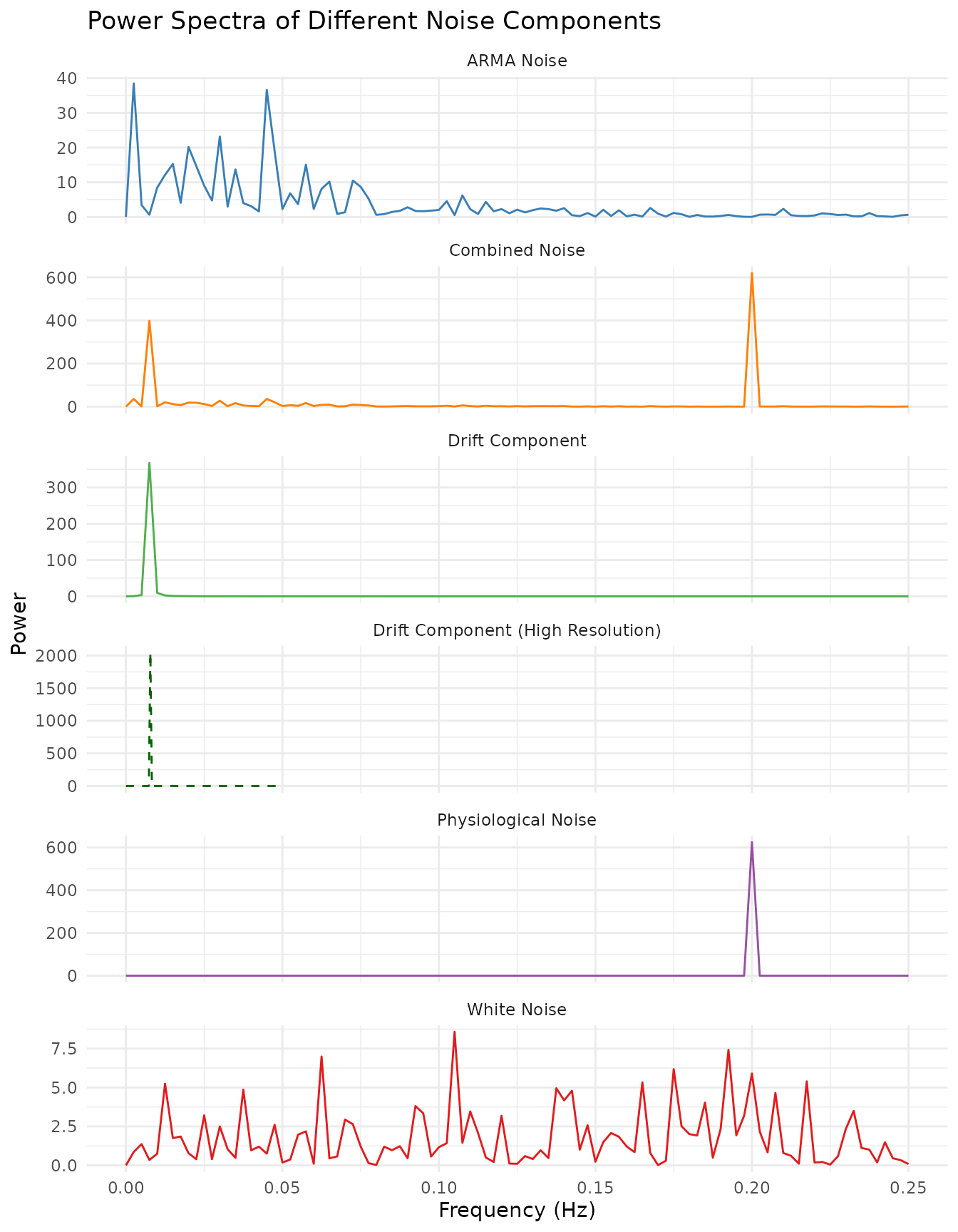
Plotting all spectra on a single log-scaled axis makes it easy to compare their relative magnitudes.
ggplot(spec_df, aes(x = Frequency, y = Power, color = NoiseType)) +
geom_line() +
theme_minimal() +
labs(title = "Power Spectra of Different Noise Components (Log Scale)",
x = "Frequency (Hz)",
y = "Power (log scale)") +
scale_y_log10() +
scale_color_brewer(palette = "Set1") +
theme(legend.position = "top") +
coord_cartesian(xlim = c(0, 0.25))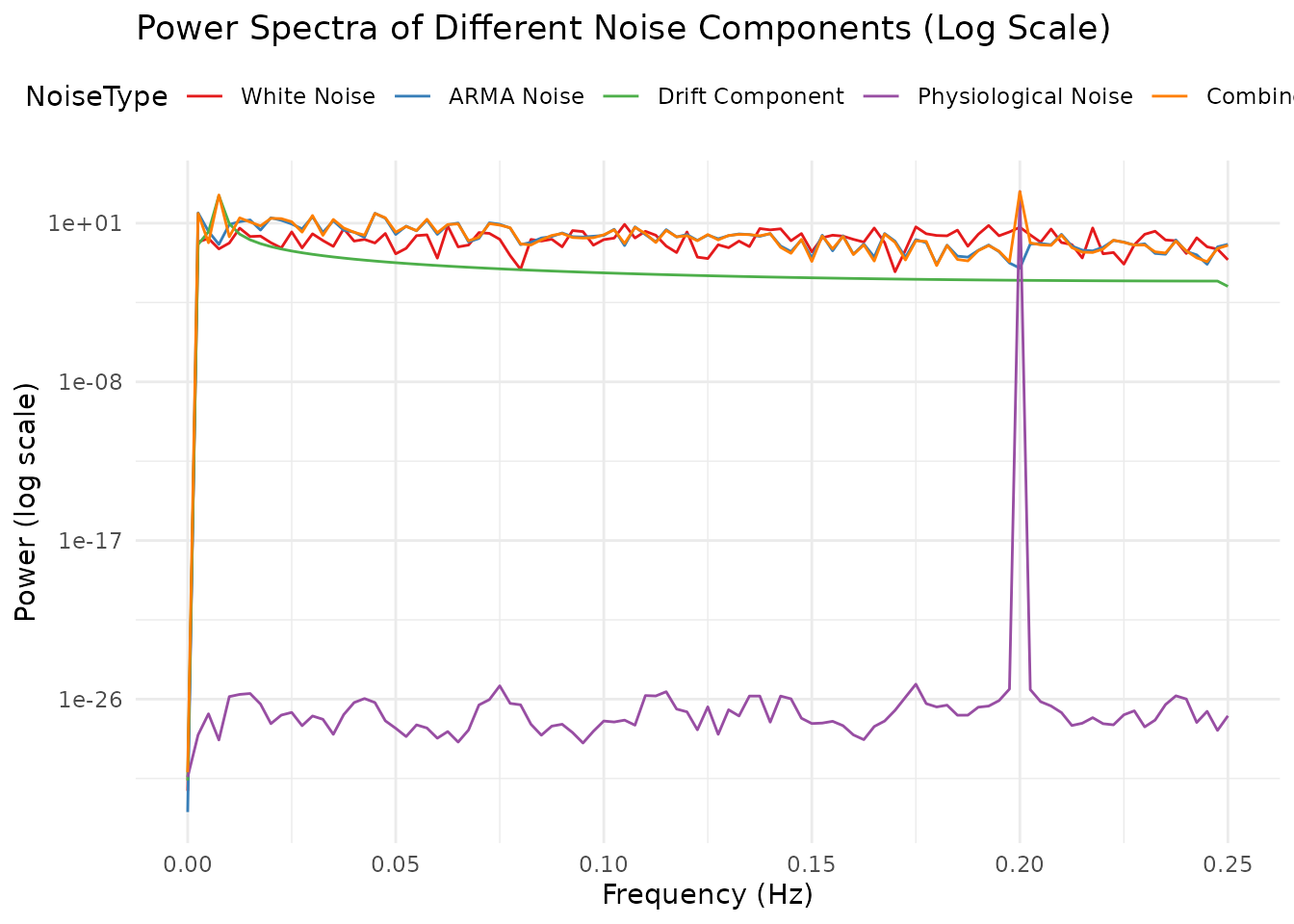
Zooming in on the very low-frequency region confirms the drift component peaks at the expected frequency (marked by the red dotted line).
drift_freq_idx <- which.min(abs(spec_df$Frequency - drift_freq))
ggplot() +
geom_line(data = subset(spec_df, NoiseType == "Drift Component"),
aes(x = Frequency, y = Power), color = "darkgreen") +
geom_line(data = subset(spec_drift_long, Frequency <= 0.05),
aes(x = Frequency, y = Power), color = "darkgreen", linetype = "dashed") +
geom_vline(xintercept = drift_freq, linetype = "dotted", color = "red") +
theme_minimal() +
labs(title = "Zoomed Power Spectrum of Drift Component",
subtitle = paste("Expected peak at", drift_freq, "Hz"),
x = "Frequency (Hz)",
y = "Power") +
coord_cartesian(xlim = c(0, 0.05))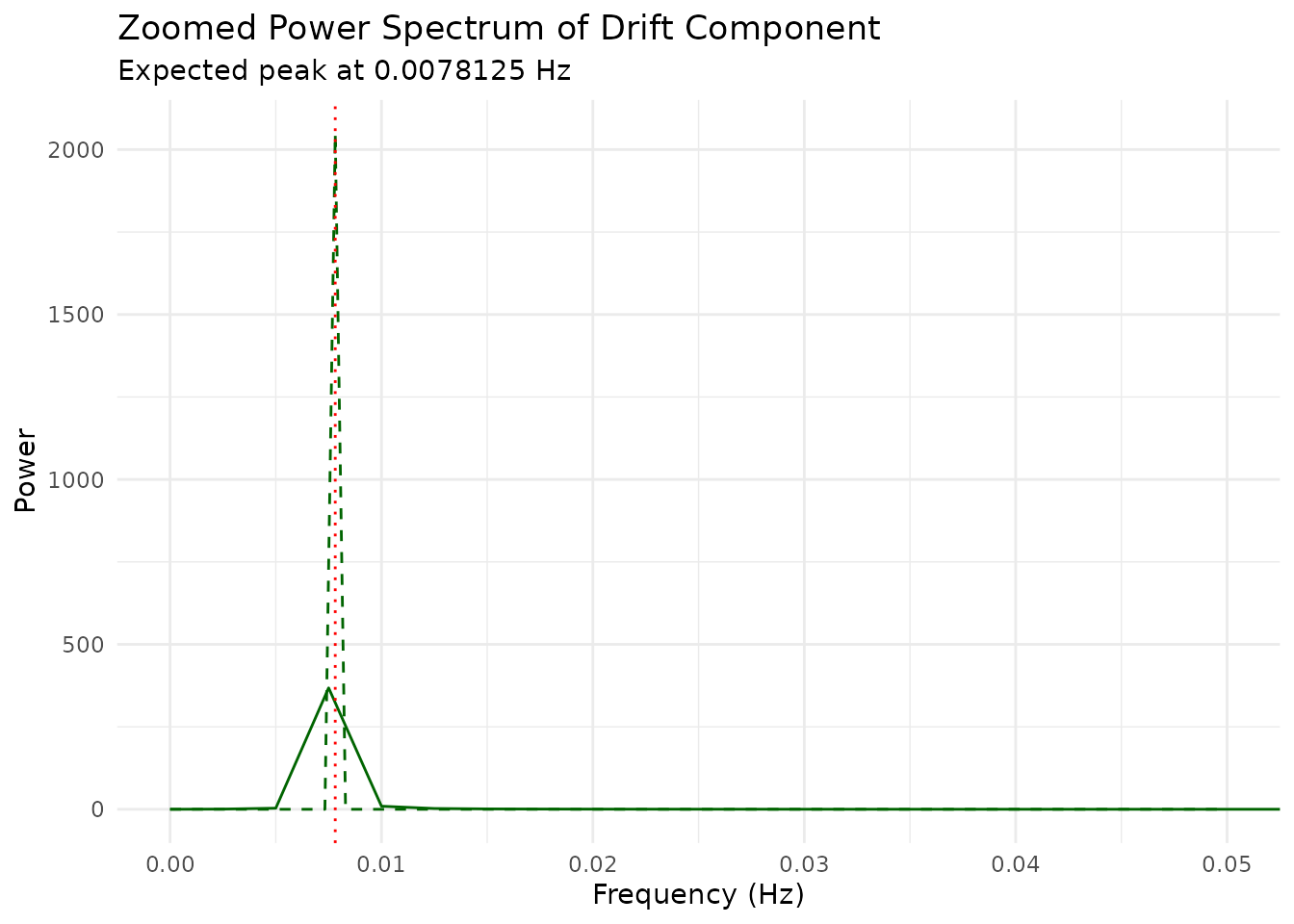
The simulation shows five distinct types of noise components:
White Noise: Random fluctuations with equal power across all frequencies (flat power spectrum).
ARMA Noise: Temporally autocorrelated noise with a characteristic “smoothed” appearance. The power spectrum shows more power at lower frequencies.
Drift Component: A very slow oscillation typical of scanner drift or physiological trends. The power spectrum shows a dominant peak at a very low frequency.
Physiological Noise: Regular oscillations at frequencies corresponding to respiration (~0.3 Hz) and cardiac-like activity (~0.8 Hz). In real fMRI data with TR=2s, cardiac frequencies (~1.2 Hz) would be aliased.
Combined Noise: All components together, creating a complex noise structure typical of real fMRI data. The power spectrum shows features from all contributing components.
These components, when added to task-related signals, create realistic fMRI time series.
Creating a Complete Dataset with Signal and Noise
The simulate_simple_dataset function combines clean
signals and noise to create a complete fMRI dataset with a specified
signal-to-noise ratio (SNR).
set.seed(42)
data_snr_1.0 <- simulate_simple_dataset(ncond = 3, TR = 2, snr = 1.0)
data_snr_0.5 <- simulate_simple_dataset(ncond = 3, TR = 2, snr = 0.5)
data_snr_0.2 <- simulate_simple_dataset(ncond = 3, TR = 2, snr = 0.2)
plot_df <- rbind(
create_plot_df(data_snr_1.0, "SNR = 1.0"),
create_plot_df(data_snr_0.5, "SNR = 0.5"),
create_plot_df(data_snr_0.2, "SNR = 0.2")
)
plot_df_long <- tidyr::pivot_longer(plot_df,
cols = -c(Time, SNR),
names_to = "Type",
values_to = "Signal")The overlay plot shows the clean signal (red) against the noisy measurement (blue) at each SNR level. As SNR decreases, the noise increasingly obscures the true signal.
ggplot(plot_df_long, aes(x = Time, y = Signal, color = Type)) +
geom_line() +
facet_wrap(~SNR, ncol = 1) +
theme_minimal() +
labs(title = "Clean vs. Noisy Signals at Different SNR Levels",
subtitle = "Single condition shown for clarity",
x = "Time (seconds)",
y = "Signal",
color = "Signal Type") +
scale_color_manual(values = c("Clean" = "darkred", "Noisy" = "steelblue")) +
theme(legend.position = "top")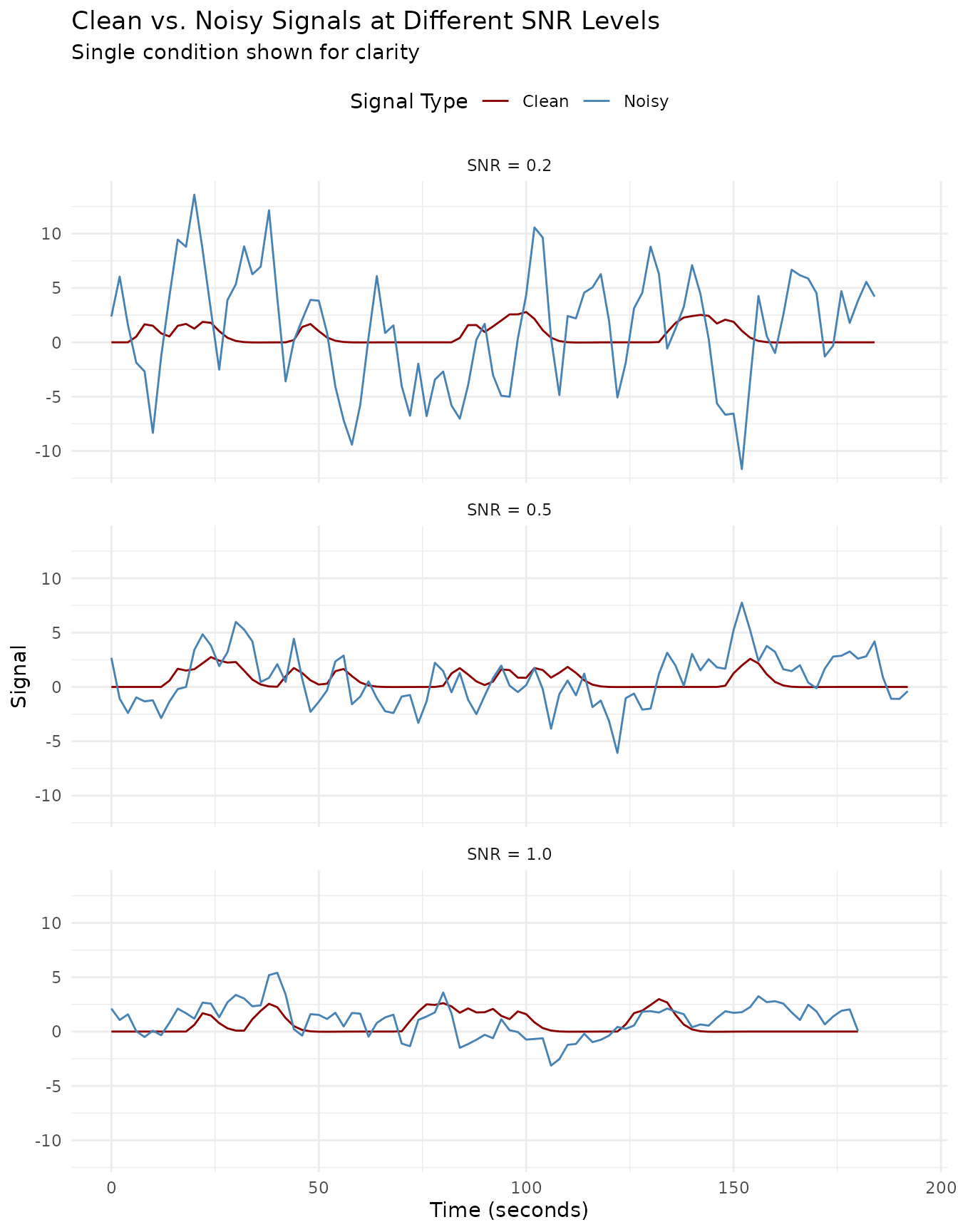
The decomposition plots below separate each SNR level into its signal, noise, and combined components. At SNR = 1.0 the signal is clearly visible; by SNR = 0.2 the noise dominates.
plot_faceted(plot_df, "SNR = 1.0")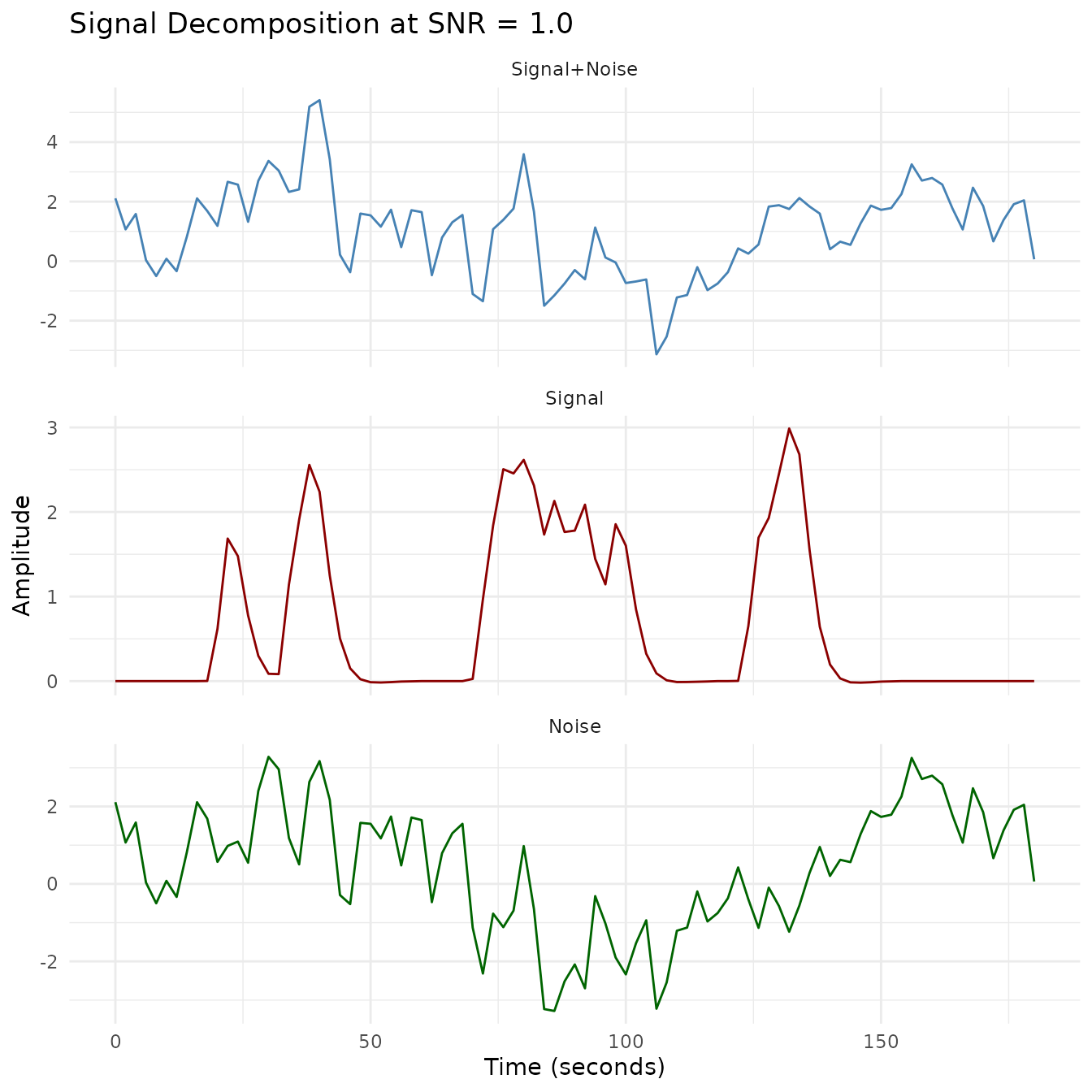
plot_faceted(plot_df, "SNR = 0.5")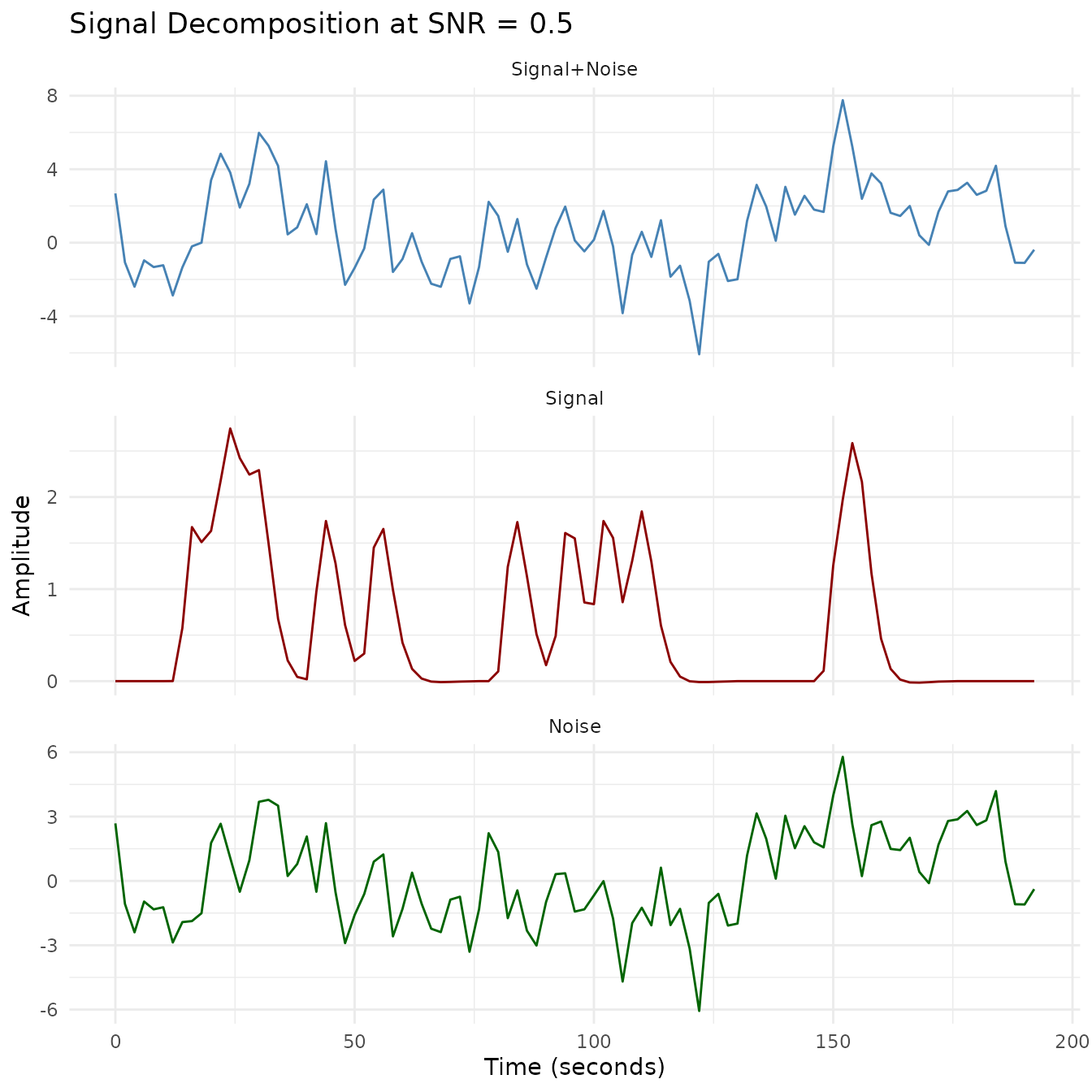
plot_faceted(plot_df, "SNR = 0.2")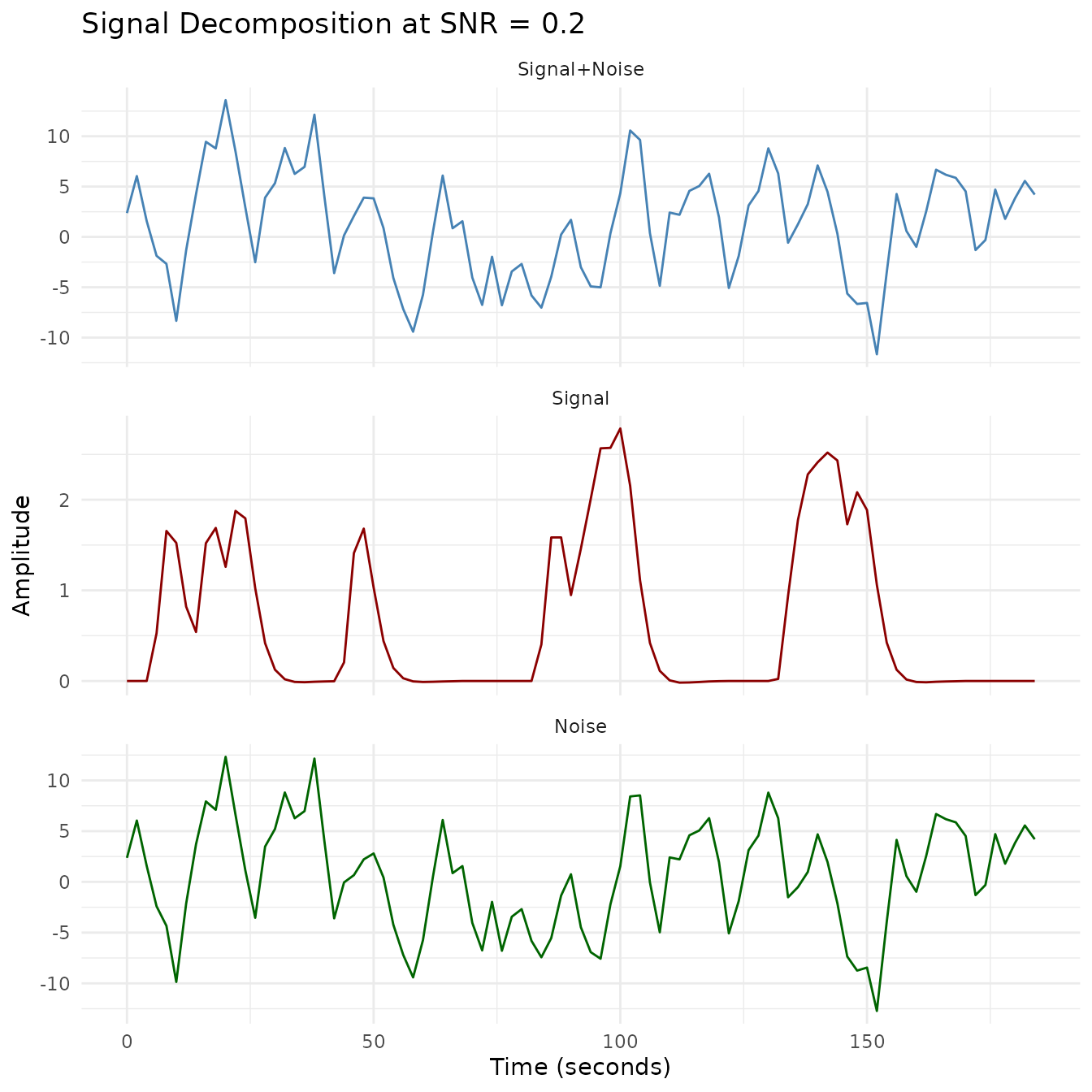
snr_stats_list <- list()
for (snr_val in unique(plot_df_long$SNR)) {
for (type_val in unique(plot_df_long$Type)) {
subset_data <- plot_df_long[plot_df_long$SNR == snr_val & plot_df_long$Type == type_val, ]
snr_stats_list[[length(snr_stats_list) + 1]] <- data.frame(
SNR = snr_val,
Type = type_val,
Mean = mean(subset_data$Signal),
SD = sd(subset_data$Signal),
Range = max(subset_data$Signal) - min(subset_data$Signal)
)
}
}
snr_stats <- do.call(rbind, snr_stats_list)
knitr::kable(snr_stats, caption = "Statistics of clean and noisy signals at different SNR levels")| SNR | Type | Mean | SD | Range |
|---|---|---|---|---|
| SNR = 1.0 | Clean | 0.6484660 | 0.9154157 | 3.005768 |
| SNR = 1.0 | Noisy | 1.0555953 | 1.5153738 | 8.541682 |
| SNR = 0.5 | Clean | 0.6082536 | 0.7916189 | 2.760946 |
| SNR = 0.5 | Noisy | 0.7358011 | 2.4147113 | 13.835178 |
| SNR = 0.2 | Clean | 0.6342752 | 0.8631817 | 2.804079 |
| SNR = 0.2 | Noisy | 1.2170041 | 5.2307652 | 25.249274 |
This visualization shows how different SNR levels affect the fMRI time series. The lower the SNR, the more the noise dominates the signal. For each SNR level, we show:
- The original clean signal (red line): The true underlying BOLD response
- The noisy signal (blue line): What would actually be measured by the scanner
- Signal decomposition: Visualization of how the signal, noise, and combined signal relate at each SNR level
With SNR = 1.0, the signal pattern remains clearly visible despite the noise. At SNR = 0.5, some features of the signal are obscured, while at SNR = 0.2, the noise substantially masks the underlying signal, making accurate detection more challenging without proper statistical methods.
The function returns: - clean: The
simulated signals without noise - noisy:
The signals with added noise - noise: The
simulated noise component - onsets: Trial
onset times - conditions: Condition labels
for each trial
Simulating Matrix Time Series with Column-Specific Variation
The most advanced simulation function,
simulate_fmri_matrix, generates multiple time series
(columns) that share the same event timing but have column-specific
variations in amplitudes and durations. This is particularly useful for
simulating multiple voxels or regions with related but slightly
different response profiles.
sim_matrix <- simulate_fmri_matrix(
n = 5, # 5 voxels/regions
total_time = 200, # 200 seconds of scan time
TR = 2, # TR = 2 seconds
n_events = 10, # 10 events
amplitudes = 1, # Base amplitude = 1
amplitude_sd = 0.3, # Amplitude variability
durations = 2, # Base duration = 2 seconds
duration_sd = 0.5, # Duration variability
noise_type = "ar1", # AR(1) noise
noise_sd = 0.5 # Noise standard deviation
)
ts_data <- sim_matrix$time_series
matrix_data <- ts_data$datamat
time_points <- seq(0, by = 2, length.out = nrow(matrix_data))
plot_data <- data.frame(Time = time_points)
for(i in 1:ncol(matrix_data)) {
plot_data[[paste0("Voxel", i)]] <- matrix_data[, i]
}
plot_data_long <- tidyr::pivot_longer(
plot_data,
cols = starts_with("Voxel"),
names_to = "Voxel",
values_to = "Signal"
)All five simulated voxels share the same event timing, but their response amplitudes and durations vary independently.
ggplot(plot_data_long, aes(x = Time, y = Signal, color = Voxel)) +
geom_line(alpha = 0.8) +
theme_minimal() +
labs(title = "Multiple Simulated Time Series with Column-Specific Variation",
x = "Time (seconds)",
y = "Signal",
color = "Voxel") +
scale_color_brewer(palette = "Set2")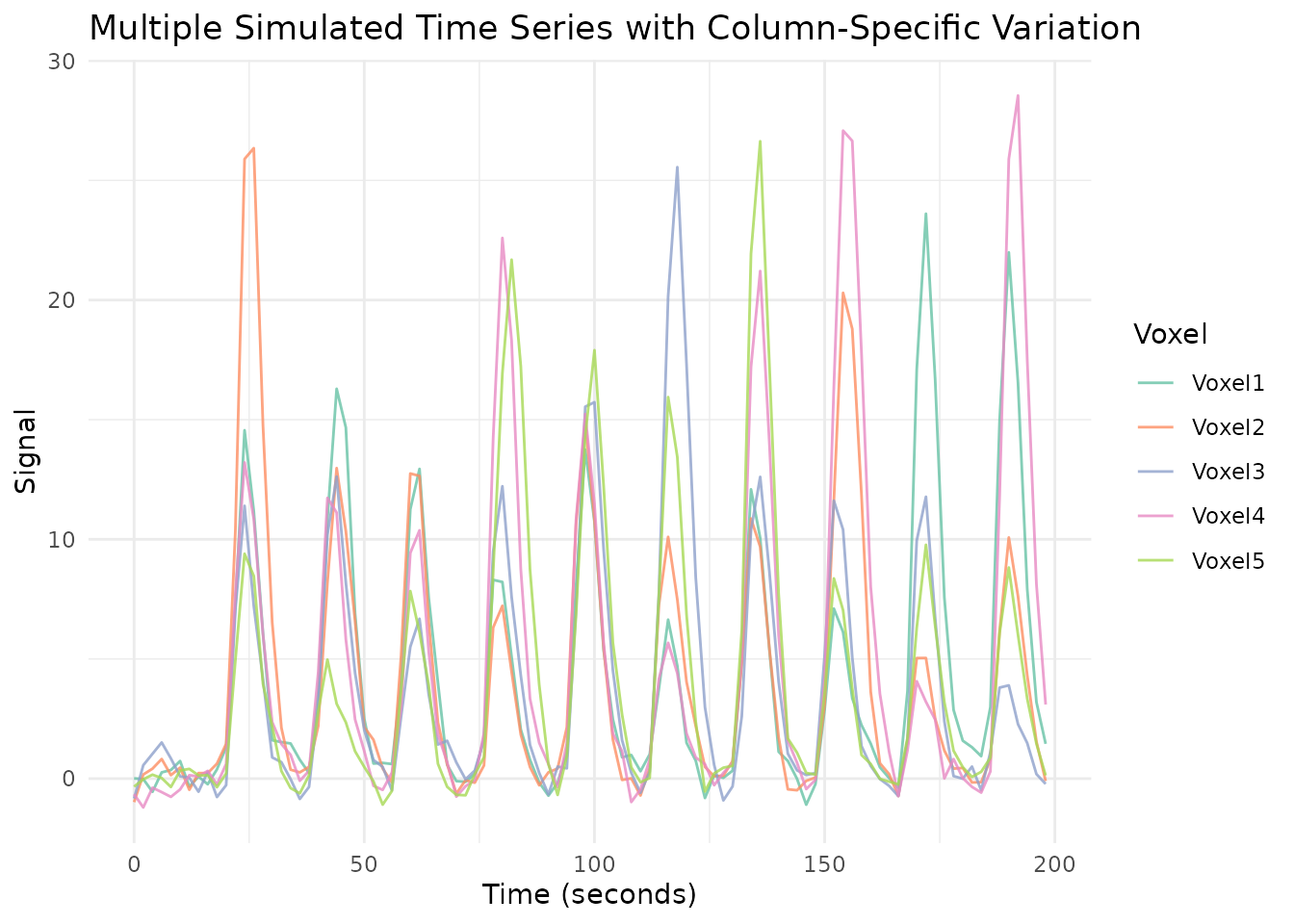
amp_df <- as.data.frame(sim_matrix$ampmat)
colnames(amp_df) <- paste0("Voxel", 1:ncol(amp_df))
amp_df$Event <- 1:nrow(amp_df)
dur_df <- as.data.frame(sim_matrix$durmat)
colnames(dur_df) <- paste0("Voxel", 1:ncol(dur_df))
dur_df$Event <- 1:nrow(dur_df)
amp_long <- tidyr::pivot_longer(
amp_df,
cols = starts_with("Voxel"),
names_to = "Voxel",
values_to = "Amplitude"
)
dur_long <- tidyr::pivot_longer(
dur_df,
cols = starts_with("Voxel"),
names_to = "Voxel",
values_to = "Duration"
)The amplitude plot shows how each voxel’s response magnitude varies from event to event around the base amplitude of 1.
ggplot(amp_long, aes(x = Event, y = Amplitude, color = Voxel, group = Voxel)) +
geom_line() +
geom_point() +
theme_minimal() +
labs(title = "Amplitude Variation Across Events and Voxels",
x = "Event Number",
y = "Amplitude",
color = "Voxel") +
scale_color_brewer(palette = "Set2")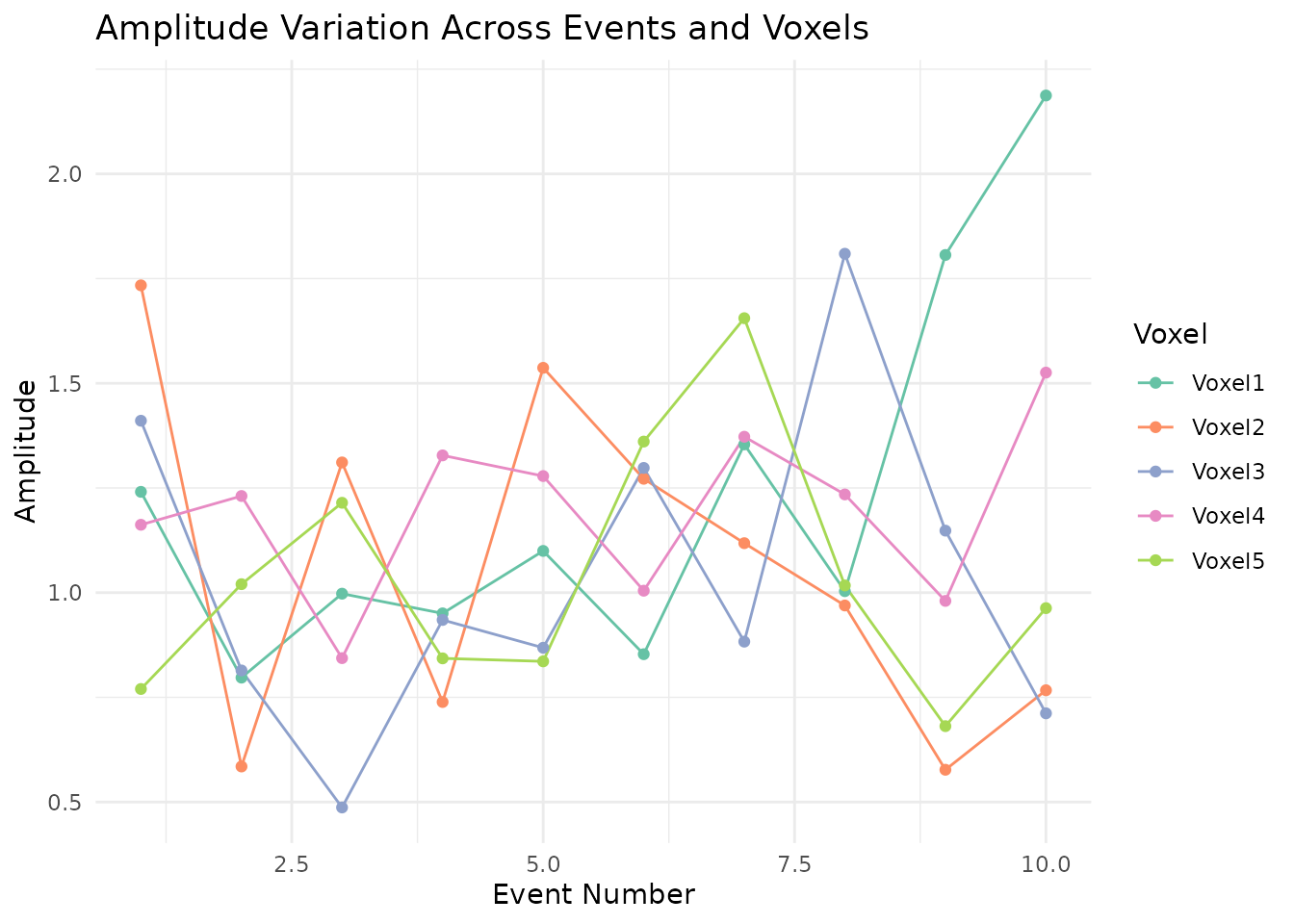
Duration variation follows the same pattern – each voxel draws its event durations independently around the base of 2 seconds.
ggplot(dur_long, aes(x = Event, y = Duration, color = Voxel, group = Voxel)) +
geom_line() +
geom_point() +
theme_minimal() +
labs(title = "Duration Variation Across Events and Voxels",
x = "Event Number",
y = "Duration (seconds)",
color = "Voxel") +
scale_color_brewer(palette = "Set2")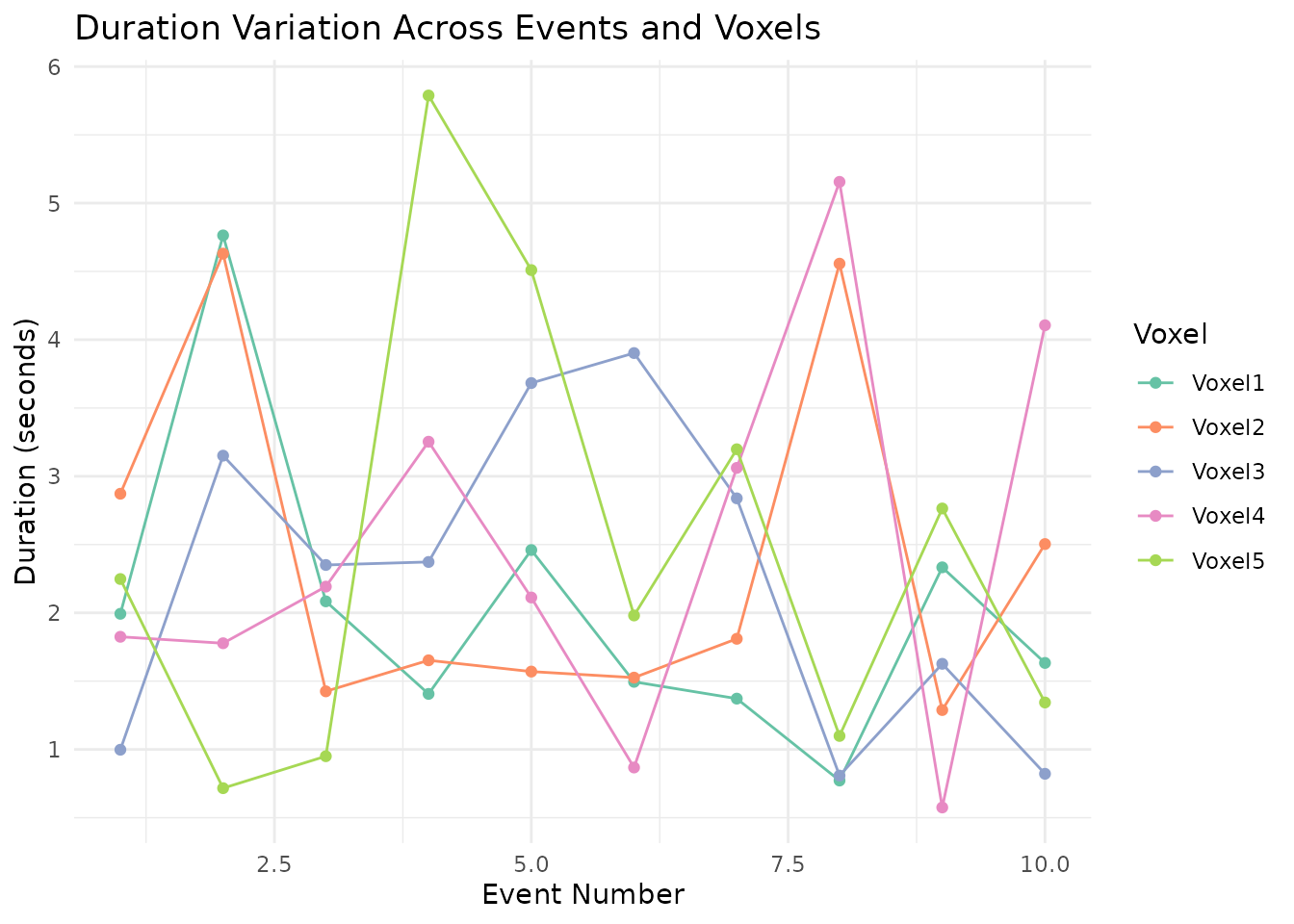
This function is particularly powerful for simulating multiple related time series with: - Shared event timing but individual variation in: - Amplitude (per event, per column) - Duration (per event, per column) - Independent noise generation for each column - Complex output including: - Time series matrix - Amplitude and duration matrices - HRF and noise parameter information
Summary and Comparison
The four simulation functions in fmrireg serve different
purposes and offer increasing levels of complexity:
-
simulate_bold_signal: Generate clean BOLD signals for multiple conditions -
simulate_noise_vector: Create realistic fMRI noise with temporal structure -
simulate_simple_dataset: Combine signal and noise with a specific SNR -
simulate_fmri_matrix: Create multiple time series with trial-by-trial, column-specific parameter variation
Choose the appropriate function based on your simulation needs: - For
basic signal generation: use simulate_bold_signal - For
realistic noise: use simulate_noise_vector - For a complete
dataset with controlled SNR: use simulate_simple_dataset -
For simulating multiple voxels/regions with shared timing but response
variation: use simulate_fmri_matrix
These functions provide a powerful toolkit for method development, validation, or teaching fMRI analysis concepts through realistic simulations.