Introduction
If you work with fMRI data, you already know the core analysis is multiple regression applied voxel-by-voxel — but specifying the design matrix, convolving events with hemodynamic response functions, and managing multi-run structures can be tedious and error-prone.
The fmrireg package gives you a concise, formula-based
interface for these tasks, inspired by lm() and
glm(). You describe your experimental design and HRF
choices in a formula, and fmrireg handles convolution, baseline
modeling, contrast computation, and model fitting.
This vignette provides a high-level overview of the key steps and
components involved in using fmrireg.
Key Components and Workflow
A typical fMRI analysis workflow with fmrireg involves
these main stages:
- Data Representation: Defining how your fMRI data and experimental design are organized.
-
Model Building: Specifying the regression model
using formulas, including:
- Defining Hemodynamic Response Functions (HRFs).
- Creating event-related regressors (
event_model). - Modeling baseline components like drift and nuisance factors
(
baseline_model). - Defining contrasts of interest (
contrast).
-
Model Estimation: Fitting the specified model to
the data (e.g., using
fmri_lm). - Advanced Topics: Exploring single-trial estimation, simulation, or translation to other packages.
1. Data Representation
Before modeling, you need to represent your fMRI data and its
structure. fmrireg uses objects like:
-
fmri_dataset/fmri_mem_dataset: Encapsulates the 4D fMRI data (potentially file-backed or in-memory), mask information, TR, run structure, and the experimental design table. -
sampling_frame: Defines the temporal structure – the number of scans per run (blocklens) and the repetition time (TR).
# Example: 5 runs, 200 scans/run, TR=2s
sframe <- sampling_frame(blocklens = rep(200, 5), TR = 2)
sframe
#> Sampling frame
#> - Blocks: 5
#> - Scans: 1000 (per block: 200, 200, 200, 200, 200 )
#> - TR: 2 s
#> - Duration: 1999 s2. Model Building
a) Hemodynamic Response Function (HRF)
The HRF models the BOLD signal response to a brief neural event.
fmrireg provides various pre-defined HRFs (e.g.,
HRF_SPMG1, HRF_GAUSSIAN) and functions to
create or modify them (gen_hrf, hrf_blocked,
hrf_lagged).
# Plot the SPM canonical HRF
time <- seq(0, 24, by = 0.2)
plot(time, HRF_SPMG1(time), type = 'l', xlab = "Time (s)", ylab = "BOLD Response",
main = "SPM Canonical HRF (HRF_SPMG1)")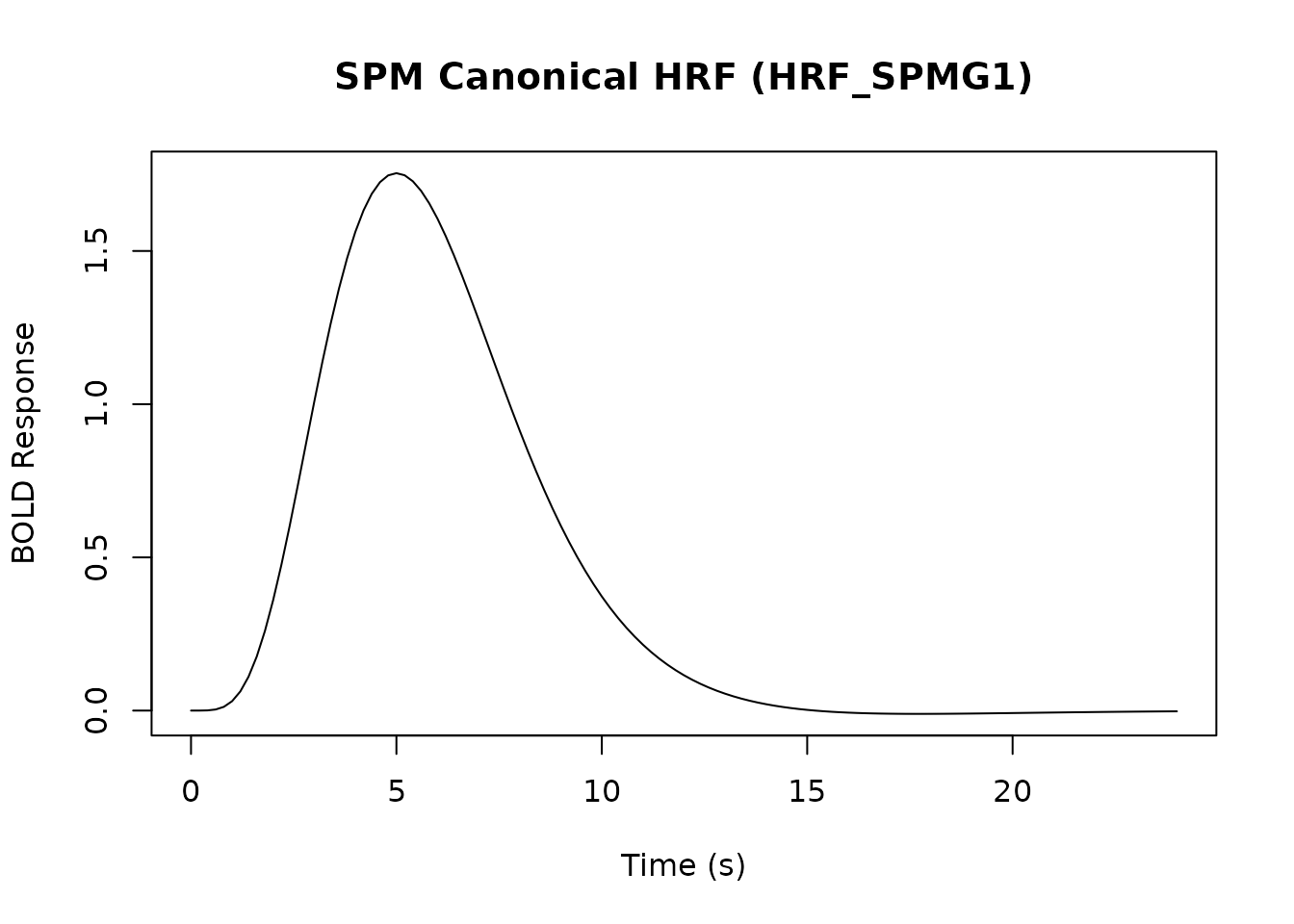
- Related Topic: Hemodynamic Response Functions
b) Event Model (event_model)
This defines the task-related part of your GLM. You specify how
experimental events (onsets, conditions, durations) map to regressors
using a formula syntax. The hrf() or
afni_hrf() functions within the formula link event
variables to specific HRF shapes.
# A simple design table with two stimulus types
design_table <- data.frame(
onset = c(10, 40, 70, 100, 130, 160, 190, 220),
stim_type = factor(rep(c("face", "scene"), 4)),
run_id = rep(1, 8)
)
# Model the effect of stim_type, convolved with the SPM canonical HRF
event_mod <- event_model(onset ~ hrf(stim_type),
data = design_table,
block = ~ run_id,
sampling_frame = sframe)
event_mod- Related Topic: Event Models
c) Baseline Model (baseline_model)
This models non-task-related components like scanner drift, run intercepts, and other nuisance factors (e.g., motion parameters).
# Example: Cubic B-spline drift model + run intercepts
# (Requires 'sframe' defined earlier)
base_mod <- baseline_model(basis = "bs", degree = 3, sframe = sframe)
print(base_mod)
#> ================================================
#> Baseline Model
#> ================================================
#> Drift Components
#> * Name: baseline_bs_3
#> * Basis type: bs
#> * Degree: 3
#> * Drift columns: 15
#>
#> Additional Components
#> * Constant columns: 5
#> * Nuisance columns: 0
#>
#> Model Summary
#> * Total columns: 20
#>
#> Design Matrix Preview
#> 0.000 0.000 0.000 0.000 ...
#> 0.015 0.000 0.000 0.000 ...
#> 0.030 0.000 0.000 0.000 ...
#> ================================================- Related Topic: Baseline Models
d) Contrasts (pair_contrast,
unit_contrast, etc.)
Contrasts define specific comparisons between condition estimates.
# Example: Contrast comparing 'face' vs 'scene' levels of 'stim_type'
face_vs_scene <- pair_contrast(~ stim_type == "face", ~ stim_type == "scene", name = "FvS")
print(face_vs_scene)
#> contrast: FvS
#> A: ~stim_type == "face"
#> B: ~stim_type == "scene"-
See Vignette:
vignette("a_05_contrasts", package = "fmrireg")
3. Model Estimation (fmri_model,
fmri_lm)
You combine the event and baseline models into a full
fmri_model and then fit it to your
fmri_dataset using estimation functions like
fmri_lm (for standard GLM) or estimate_betas
(for single-trial betas).
# Combine event and baseline models
full_model <- fmri_model(event_mod, base_mod)
# Fit the GLM
# (Requires 'my_dataset' of class fmri_dataset)
fit <- fmri_lm(full_model, dataset = my_dataset)
# Robust fit down-weighting outlier time points
fit_robust <- fmri_lm(full_model, dataset = my_dataset, robust = TRUE)
# Extract contrast results (if contrasts were defined)
# con_results <- coef(fit) # Get coefficientsSetting robust = TRUE invokes row-wise weighting to
lessen the influence of motion spikes or other frame-level
artifacts.
-
See Vignettes:
vignette("a_09_linear_model", package = "fmrireg")- Linear Models and Trialwise Analysis
4. Advanced Topics
fmrireg also supports:
-
Simulation: Creating artificial fMRI data
(
simulate_fmri_matrix). -
AFNI Translation: Generating scripts for AFNI’s
3dDeconvolve(afni_lm). -
Low-level Regressors: Manually creating and
manipulating regressors (
regressor).
Acceleration: Sketched / Nyström Analysis
For large problems, you can accelerate voxelwise GLM estimation with: - SRHT/IHS temporal sketching to reduce timepoints while preserving LS geometry. - Nyström/landmark spatial extension to solve on a subset of voxels and extend with parcel‑aware kernels.
See vignette("sketched-ar", package = "fmrireg") for
guidance, tradeoffs, and practical defaults.
Group-Level Meta-Analysis
Combine subject‑level maps or ROI summaries using
fixed/random‑effects meta‑regression: - Inputs: HDF5/NIfTI (beta+SE or
t+df) or ROI CSV. - Random‑effects: Paule–Mandel/DerSimonian–Laird/REML
alias; optional robust Huber. - t‑only combine:
Stouffer/Fisher/Lancaster (supports equal/IVW/custom weights where
applicable). - Exact contrasts: at fit‑time (contrasts=…) or post‑hoc
via return_cov = "tri" + contrast(). -
Multiple comparisons: BH FDR or spatial FDR (block/parcel aware).
See vignette("group_analysis", package = "fmrireg") for
end‑to‑end examples.
Getting Started
This overview provides a map of the fmrireg package. For
detailed explanations and examples, please refer to the specific
vignettes linked above.
Next Steps
-
vignette("a_10_dataset", package = "fmrireg")— Dataset management -
vignette("a_08_simulation", package = "fmrireg")— Simulating fMRI data -
vignette("group_analysis", package = "fmrireg")— Group analysis -
vignette("sketched-ar", package = "fmrireg")— Sketched/Nyström acceleration