Group Analysis
Bradley R. Buchsbaum
2026-04-15
Source:vignettes/group_analysis.Rmd
group_analysis.RmdOverview
This vignette walks through a compact, end-to-end example of group analysis with fmrireg. We first construct a small ROI-level dataset to illustrate the basic meta-regression interface, and then move to a voxelwise example using tiny synthetic NIfTI files. Along the way we show how to compare groups with fixed- and random-effects meta-analysis, how to obtain exact contrasts either at fit-time or post-hoc by saving covariance, and how to perform group inference from t-maps alone using Stouffer, Fisher, or Lancaster combinations.
Create a small ROI dataset
We simulate 10 subjects across two groups (A/B) for a single ROI. Group B has an effect that is 1 unit larger than group A. All subjects have the same SE for clarity.
n_per_group <- 5
subjects <- sprintf("s%02d", 1:(2 * n_per_group))
group <- factor(rep(c("A", "B"), each = n_per_group))
beta <- ifelse(group == "A", 0.5, 1.5)
se <- rep(0.25, length(beta))
roi_df <- data.frame(
subject = subjects,
roi = "ExampleROI",
beta = beta,
se = se,
group = group,
stringsAsFactors = FALSE
)
gd <- group_data_from_csv(
roi_df,
effect_cols = c(beta = "beta", se = "se"),
subject_col = "subject",
roi_col = "roi",
covariate_cols = c("group")
)
gd
#> Group Data Object
#> Format: csv
#> Subjects: 10
#> Covariates: groupFit group meta-analysis
We first fit an intercept-only model, then a model including a group term.
fit_fe <- fmri_meta(gd, formula = ~ 1, method = "fe", verbose = FALSE)
fit_cov <- fmri_meta(gd, formula = ~ 1 + group, method = "fe", verbose = FALSE)
print(fit_cov)
#> fMRI Meta-Analysis Results
#> ==========================
#>
#> Method: fe
#> Robust: none
#> Formula: ~1 + group
#> Subjects: 10
#> ROIs analyzed: 1
#>
#> Heterogeneity:
#> Mean tau^2: 0
#> Mean I^2: NaN %
summary(fit_cov)
#> fMRI Meta-Analysis Summary
#> ==========================
#>
#> fMRI Meta-Analysis Results
#> ==========================
#>
#> Method: fe
#> Robust: none
#> Formula: ~1 + group
#> Subjects: 10
#> ROIs analyzed: 1
#>
#> Heterogeneity:
#> Mean tau^2: 0
#> Mean I^2: NaN %
#>
#> Coefficients:
#> (Intercept):
#> Mean effect: 0.5
#> Mean SE: 0.1118034
#> Significant:1/1 (100%)
#> groupB:
#> Mean effect: 1
#> Mean SE: 0.1581139
#> Significant:1/1 (100%)Note: With equal SE per subject, fixed-effects and random-effects
will yield similar point estimates. Random-effects
(method = "pm") will estimate between- subject
heterogeneity (tau2) when present.
fit_pm <- fmri_meta(gd, formula = ~ 1 + group, method = "pm", verbose = FALSE)
summary(fit_pm)
#> fMRI Meta-Analysis Summary
#> ==========================
#>
#> fMRI Meta-Analysis Results
#> ==========================
#>
#> Method: pm
#> Robust: none
#> Formula: ~1 + group
#> Subjects: 10
#> ROIs analyzed: 1
#>
#> Heterogeneity:
#> Mean tau^2: 0
#> Mean I^2: NaN %
#>
#> Coefficients:
#> (Intercept):
#> Mean effect: 0.5
#> Mean SE: 0.1118034
#> Significant:1/1 (100%)
#> groupB:
#> Mean effect: 1
#> Mean SE: 0.1581139
#> Significant:1/1 (100%)Extract coefficients and a contrast
coef_names <- colnames(fit_cov$coefficients)
coef_names
#> [1] "(Intercept)" "groupB"
coef_est <- as.numeric(fit_cov$coefficients[1, ])
names(coef_est) <- coef_names
coef_est
#> (Intercept) groupB
#> 0.5 1.0
if (any(grepl("group", coef_names))) {
w <- rep(0, length(coef_names)); names(w) <- coef_names
w[grep("group", coef_names)] <- 1
con <- contrast(fit_cov, w)
con
}
#> $estimate
#> [1] 1
#>
#> $se
#> [1] 0.1581139
#>
#> $z
#> [1] 6.324555
#>
#> $p
#> [,1]
#> ExampleROI 2.539629e-10
#>
#> $weights
#> (Intercept) groupB
#> 0 1
#>
#> $name
#> [1] "groupB"
#>
#> $parent
#> fMRI Meta-Analysis Results
#> ==========================
#>
#> Method: fe
#> Robust: none
#> Formula: ~1 + group
#> Subjects: 10
#> ROIs analyzed: 1
#>
#> Heterogeneity:
#> Mean tau^2: 0
#> Mean I^2: NaN %
#>
#> attr(,"class")
#> [1] "fmri_meta_contrast"A quick visualization
We can visualize the group effects and their 95% CIs for the ROI-level fit.
df_tidy <- tidy(fit_cov, conf.int = TRUE)
df_tidy
#> # A tibble: 2 × 10
#> roi term estimate std.error statistic p.value tau2 I2 conf.low
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 ExampleROI (Interc… 0.5 0.112 4.47 7.74e- 6 0 NA 0.281
#> 2 ExampleROI groupB 1 0.158 6.32 2.54e-10 0 NA 0.690
#> # ℹ 1 more variable: conf.high <dbl>
ggplot(df_tidy, aes(x = term, y = estimate, ymin = conf.low, ymax = conf.high)) +
geom_pointrange() +
geom_hline(yintercept = 0, linetype = 2) +
labs(title = "ROI-level group meta-analysis",
x = "Term", y = "Estimate +/- 95% CI") +
theme_minimal()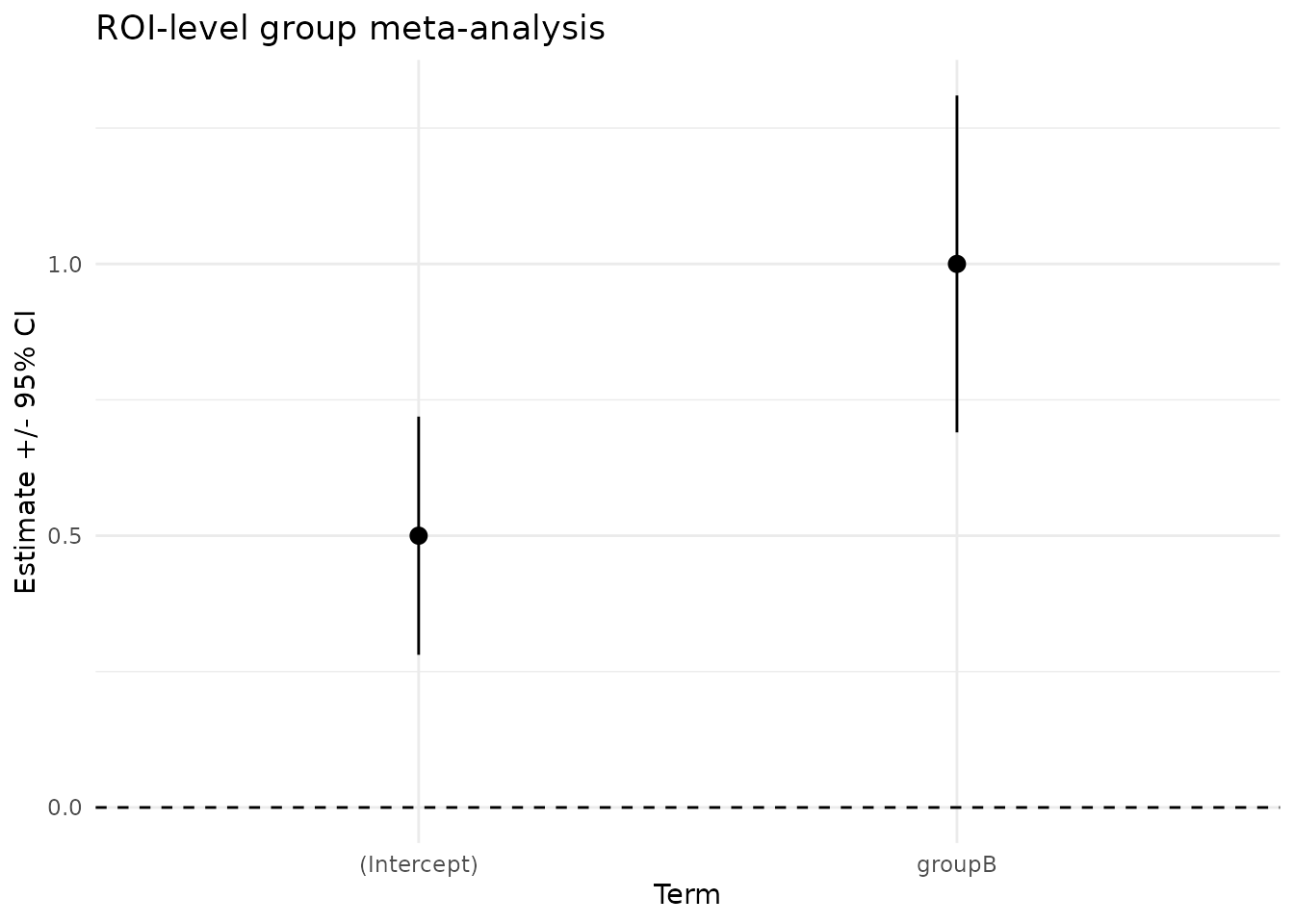
Notes on voxelwise analysis
For voxelwise analysis, construct group_data with format
"nifti" or "h5":
# HDF5 (produced via write_results.fmri_lm)
# gd_h5 <- fmrireg::group_data(h5_paths, format = "h5",
# subjects = subject_ids,
# covariates = covariates_df,
# contrast = "ContrastName")
# NIfTI (provide per-subject paths for beta/SE or t)
# gd_nii <- fmrireg::group_data(list(beta = beta_paths, se = se_paths), format = "nifti",
# subjects = subject_ids,
# mask = "group_mask.nii.gz")
# fit <- fmri_meta(gd_h5, formula = ~ 1 + group, method = "pm")
# fit <- fmri_meta(gd_nii, formula = ~ 1, method = "fe")For multiple comparisons correction that leverages spatial structure,
see spatial_fdr() and create_3d_blocks().
Minimal NIfTI Example (Reproducible)
This chunk creates tiny synthetic NIfTI volumes on disk (in a temp dir) for a voxelwise demonstration. Group B has a higher effect in a small cube.
library(neuroim2)
set.seed(42)
tmpdir <- tempdir()
space <- NeuroSpace(c(8, 8, 8), spacing = c(2, 2, 2))
n_per_group <- 3
ids <- sprintf("sub-%02d", 1:(2 * n_per_group))
grp <- factor(rep(c("A", "B"), each = n_per_group))
active <- array(FALSE, dim = c(8, 8, 8))
active[3:5, 3:5, 3:5] <- TRUE
beta_paths <- character(length(ids))
se_paths <- character(length(ids))
for (i in seq_along(ids)) {
b <- array(0, dim = c(8, 8, 8))
b[active] <- if (grp[i] == "A") 0.5 else 1.5
b <- b + array(rnorm(length(b), sd = 0.05), dim = dim(b))
v_beta <- NeuroVol(b, space)
s <- array(0.25, dim = c(8, 8, 8))
v_se <- NeuroVol(s, space)
beta_paths[i] <- file.path(tmpdir, sprintf("%s_beta.nii.gz", ids[i]))
se_paths[i] <- file.path(tmpdir, sprintf("%s_se.nii.gz", ids[i]))
write_vol(v_beta, beta_paths[i])
write_vol(v_se, se_paths[i])
}
mask_path <- file.path(tmpdir, "mask.nii.gz")
write_vol(NeuroVol(array(1, dim = c(8, 8, 8)), space), mask_path)
gd_nii <- group_data_from_nifti(
beta_paths = beta_paths,
se_paths = se_paths,
subjects = ids,
covariates = data.frame(group = grp),
mask = mask_path
)
fit_nii <- fmri_meta(gd_nii, formula = ~ 1 + group, method = "fe", verbose = FALSE)
fit_nii
#> fMRI Meta-Analysis Results
#> ==========================
#>
#> Method: fe
#> Robust: none
#> Formula: ~1 + group
#> Subjects: 6
#> Voxels analyzed: 512
#>
#> Heterogeneity:
#> Mean tau^2: 0
#> Mean I^2: 0 %
group_col <- grep("group", colnames(fit_nii$coefficients))
pvals <- 2 * pnorm(-abs(fit_nii$coefficients[, group_col] / fit_nii$se[, group_col]))
sum(pvals < 0.05)
#> [1] 27
sfr <- spatial_fdr(fit_nii, p = colnames(fit_nii$coefficients)[group_col], group = "blocks")
sum(sfr$reject)
#> [1] 75
img_group_est <- coef_image(fit_nii, colnames(fit_nii$coefficients)[group_col], statistic = "estimate")
range(as.array(img_group_est), na.rm = TRUE)
#> [1] -0.09585902 1.08515382Exact contrasts and stored covariance
You can request exact contrasts at fit-time or store per-voxel covariance for exact post-hoc contrasts.
fit_nii_pm <- fmri_meta(
gd_nii, formula = ~ 1 + group, method = "pm",
return_cov = "tri", verbose = FALSE
)
con <- contrast(fit_nii_pm, c("(Intercept)" = 0, group = 1))
summary(con)
#> Length Class Mode
#> estimate 512 -none- numeric
#> se 512 -none- numeric
#> z 512 -none- numeric
#> p 512 -none- numeric
#> weights 2 -none- numeric
#> name 1 -none- character
#> parent 17 fmri_meta list
fit_nii_con <- fmri_meta(
gd_nii, formula = ~ 1 + group, method = "pm",
contrasts = matrix(c(0, 1), nrow = 1,
dimnames = list("group", colnames(fit_nii_pm$model$X))),
verbose = FALSE
)
str(fit_nii_con$contrasts)
#> List of 4
#> $ names : chr "group"
#> $ estimate: num [1:512, 1] 2.05e-02 1.79e-02 -3.49e-03 -5.51e-02 3.53e-05 ...
#> $ se : num [1:512, 1] 0.204 0.204 0.204 0.204 0.204 ...
#> $ z : num [1:512, 1] 0.100565 0.0878 -0.017112 -0.269929 0.000173 ...Two-sample t-test (Welch and OLS) on NIfTI
As an alternative to meta-analysis, we can run two-sample voxelwise t-tests directly on the per-subject beta maps, using either Welch’s unequal-variance test or a standard OLS/Student t-test via a simple design matrix.
fit_welch <- fmri_ttest(gd_nii, formula = ~ 1 + group, engine = "welch")
t_welch <- as.numeric(fit_welch$t["group", ])
df_welch <- as.numeric(fit_welch$df["group", ])
p_welch <- 2 * pt(abs(t_welch), df = df_welch, lower.tail = FALSE)
fit_ols <- fmri_ttest(gd_nii, formula = ~ 1 + group, engine = "classic")
rn_t <- rownames(fit_ols$t)
if (!is.null(rn_t) && any(rn_t == "group")) {
t_ols <- fit_ols$t["group", ]
df_ols <- as.numeric(fit_ols$df["group", ])
} else {
t_ols <- fit_ols$t[2, ]
df_ols <- rep(fit_ols$df[2, 1], length(t_ols))
}
p_ols <- 2 * pt(abs(t_ols), df = df_ols, lower.tail = FALSE)
timg_welch <- NeuroVol(array(NA_real_, dim = c(8, 8, 8)), space)
mask_img <- if (!is.null(gd_nii$mask_data)) gd_nii$mask_data else neuroim2::read_vol(mask_path)
timg_welch[as.array(mask_img) > 0] <- t_welch
range(as.array(timg_welch), na.rm = TRUE)
#> [1] -81.539233 4.582865
sum(p_welch < 0.05)
#> [1] 46
sum(p_ols < 0.05)
#> [1] 51Combining t-statistics only (Stouffer/Fisher/Lancaster)
When only per-subject t-statistics and degrees-of-freedom are
available, you can still carry out group inference without betas/SEs by
setting combine = in fmri_meta() (or in
fmri_ttest(..., engine = "meta")). Stouffer combines signed
z-scores and supports equal, inverse-variance or custom weights; Fisher
combines p-values with equal weights; and Lancaster provides a weighted
Fisher variant by mapping weights to per-subject degrees-of-freedom.
dat_full <- read_nifti_full(gd_nii)
tmat <- dat_full$beta / dat_full$se
t_paths <- character(length(ids))
for (i in seq_along(ids)) {
img <- NeuroVol(array(NA_real_, dim = c(8, 8, 8)), space)
img[as.array(neuroim2::read_vol(mask_path)) > 0] <- tmat[i, ]
pth <- file.path(tmpdir, sprintf("%s_t.nii.gz", ids[i]))
write_vol(img, pth)
t_paths[i] <- pth
}
gd_t <- group_data_from_nifti(
t_paths = t_paths,
df = 60,
subjects = ids,
covariates = data.frame(group = grp),
mask = mask_path
)
fit_st <- fmri_meta(gd_t, formula = ~ 1, combine = "stouffer", verbose = FALSE)
w_subj <- rep(1, length(ids))
fit_st_w <- fmri_meta(gd_t, formula = ~ 1, combine = "stouffer",
weights = "custom", weights_custom = w_subj,
verbose = FALSE)
fit_fi <- fmri_meta(gd_t, formula = ~ 1, combine = "fisher", verbose = FALSE)
fit_la <- fmri_meta(gd_t, formula = ~ 1, combine = "lancaster",
weights = "custom", weights_custom = w_subj,
verbose = FALSE)Meta engine via fmri_ttest with weights
The t-test interface supports the meta engine with equal/custom weighting.
fit_tt_meta <- fmri_ttest(gd_nii, formula = ~ 1 + group, engine = "meta",
weights = "equal")
w_subj <- rep(1, length(ids))
fit_tt_meta_w <- fmri_ttest(gd_nii, formula = ~ 1 + group, engine = "meta",
weights = "custom", weights_custom = w_subj)
fit_tt_la <- fmri_ttest(gd_t, formula = ~ 1, engine = "meta",
combine = "lancaster", weights = "custom",
weights_custom = w_subj)ROI t-only example (Stouffer and Lancaster)
You can also combine t-statistics at the ROI level from a tabular CSV. Provide per-subject t and df, then choose a combine method.
roi_t_df <- data.frame(
subject = subjects,
roi = "ExampleROI",
t = rnorm(length(subjects), mean = 2.0, sd = 0.5),
df = 40,
stringsAsFactors = FALSE
)
gd_roi_t <- group_data_from_csv(
roi_t_df,
effect_cols = c(t = "t", df = "df"),
subject_col = "subject",
roi_col = "roi"
)
fit_roi_st <- fmri_meta(gd_roi_t, formula = ~ 1, combine = "stouffer", verbose = FALSE)
w_roi <- rep(1, length(subjects))
fit_roi_la <- fmri_meta(gd_roi_t, formula = ~ 1, combine = "lancaster",
weights = "custom", weights_custom = w_roi,
verbose = FALSE)
c(fit_roi_st$method, fit_roi_la$method)
#> [1] "pm" "pm"A brief recap
Meta-analysis in fmrireg supports fixed-effects and several
random-effects estimators (method = "fe"|"pm"|"dl"|"reml"),
with optional robust Huber weighting. You can pass subject-level
covariates for group comparisons and, when working from t-maps only, set
combine = to use Stouffer, Fisher, or Lancaster. Exact
contrasts are available either at fit-time (via contrasts=)
or post-hoc by saving packed covariance with
return_cov = "tri" and then calling
contrast(). Weighting applies to both meta-regression and
t-only combinations (weights = "ivw"|"equal"|"custom", with
weights_custom as a vector of length subjects or an S x P
matrix). The examples above show ROI-based meta-regression, voxelwise
fits from NIfTI, and t-only combinations via both
fmri_meta() and
fmri_ttest(..., engine = "meta").