Surface Parcellations with neurosurf
neuroatlas Dev Team
2026-02-26
Source:vignettes/surface-parcellations.Rmd
surface-parcellations.RmdWhat this vignette covers
This guide focuses on loading surface parcellations
as actual meshes (neurosurf::LabeledNeuroSurface)—not ggseg
flats. We cover:
- Discovering available surface templates in TemplateFlow
- Schaefer on fsaverage6 (packaged geometry)
- Schaefer on fsaverage5/fsaverage (via TemplateFlow, when available)
- Glasser MMP1.0 (via TemplateFlow, when available)
- Loading raw surface geometry from TemplateFlow
- Downloading surfaces to a local folder
- What files are fetched, what the
dataslot contains, and how labels map. - Minimal patterns to attach your own per-vertex values.
Discovering available surface templates
Before loading surfaces, you can query what’s available in TemplateFlow:
# Find surface-related template spaces
tflow_spaces(pattern = "fs")
#> [1] "fsLR" "fsaverage"
# List available fsLR surfaces
tflow_files("fsLR", query_args = list(hemi = "L"))
# Filter by specific density and surface type
tflow_files("fsLR", query_args = list(
hemi = "L",
density = "32k",
suffix = "midthickness"
))
# List fsaverage surfaces at a specific resolution (e.g., fsaverage6)
tflow_files("fsaverage", query_args = list(
hemi = "L",
resolution = "06" # fsaverage6
))Common query parameters for surfaces: - hemi: “L” or “R”
- density: “32k”, “164k” (for fsLR) -
resolution: “06” (fsaverage6), “05” (fsaverage5) -
suffix: “pial”, “white”, “inflated”, “midthickness”,
“sphere”
Schaefer surface atlases
# 200 parcels, 7 networks, fsaverage6 inflated (packaged geometry, no TF needed)
atl <- schaefer_surf(
parcels = 200,
networks = 7,
space = "fsaverage6",
surf = "inflated"
)Returned structure:
-
atl$lh_atlas,atl$rh_atlas:LabeledNeuroSurfaceobjects. -
slot(..., "data"): integer parcel IDs per vertex. -
atl$labels/atl$orig_labels: name lookups;atl$network: network per ID.
Using TemplateFlow spaces (when available)
Note: TemplateFlow surface support requires the
fsaverage template to be available in your TemplateFlow
installation. As of this writing, surface templates may have limited
availability. Use fsaverage6 (above) for reliable access to
Schaefer atlases.
# fsaverage5 white surface; uses TemplateFlow geometry (if available)
atl_tf <- schaefer_surf(
parcels = 400,
networks = 17,
space = "fsaverage5",
surf = "white"
)If TemplateFlow surfaces are available, this uses the same labels/data layout as above.
Attaching your own values
vals <- rnorm(length(atl$ids)) # one value per parcel
mapped <- map_atlas(atl, vals) # returns LabeledNeuroSurface per hemi with data filledmap_atlas() writes your parcel values into the
data slot of each hemisphere surface.
Glasser MMP1.0 surface atlas (when available)
Note: Like Schaefer TemplateFlow surfaces, Glasser surface support
requires the fsaverage template in TemplateFlow. Check
availability before using.
# Requires fsaverage template in TemplateFlow
glas <- glasser_surf(space = "fsaverage", surf = "pial")
slot(glas$lh_atlas, "data")[1:5] # parcel IDs
glas$labels[1:5] # names for those IDsIf available, geometry comes from TemplateFlow (fsaverage
pial/white/inflated/midthickness); labels come from the HCP-MMP1.0
fsaverage annotations published on Figshare as the “HCP-MMP1_0 projected
on fsaverage” dataset (ID 3498446; DOI: 10.6084/m9.figshare.3498446).
The data vector length equals the vertex count of the mesh;
each entry is the parcel ID at that vertex.
What’s in the data slot?
- Schaefer / Glasser surface atlases: integer parcel IDs per vertex.
- If you construct a
NeuroSurfaceyourself (e.g., fromload_surface_template()), you supply the per-vertex numeric values.
Saving / reusing
saveRDS(atl, "schaefer200_7_fs6_inflated.rds")
# reload later
atl2 <- readRDS("schaefer200_7_fs6_inflated.rds")Sanity check: parcel surface renders
We snapshot a labeled Schaefer surface to verify geometry+labels load. Runs only when TemplateFlow is available (for non-packaged spaces).
dir.create("figures", showWarnings = FALSE)
png_path <- file.path("figures", "schaefer200_fs6_L_inflated.png")
atl <- schaefer_surf(
parcels = 200,
networks = 7,
space = "fsaverage6", # packaged geometry (no TF), keeps it fast
surf = "inflated"
)
geom_l <- neurosurf::geometry(atl$lh_atlas)
neurosurf::snapshot_surface(geom_l, file = png_path)
png_pathIf the image renders, the parcellated surface is displayable.
Loading raw surface geometry from TemplateFlow
To load just the surface geometry (without parcellation labels), use
get_surface_template() or
load_surface_template():
# Get path to a surface file
fslr_path <- get_surface_template(
template_id = "fsLR",
surface_type = "midthickness",
hemi = "L",
density = "32k"
)
print(fslr_path)
# Load as a neurosurf SurfaceGeometry object
fslr_geom <- load_surface_template(
template_id = "fsLR",
surface_type = "inflated",
hemi = "L",
density = "32k"
)
# Load both hemispheres at once
both_hemi <- load_surface_template(
template_id = "fsLR",
surface_type = "pial",
hemi = "both",
density = "32k"
)
# Returns list with $L and $RDownloading surfaces to a local folder
To copy TemplateFlow surfaces to a local directory:
# Get paths to surfaces (automatically downloaded to cache)
files <- tflow_files("fsLR", query_args = list(hemi = "L", density = "32k"))
# Copy to local folder
dest_folder <- "~/my_surfaces/fsLR"
dir.create(dest_folder, recursive = TRUE, showWarnings = FALSE)
file.copy(files, dest_folder)
# Or use a helper function for bulk downloads
download_surfaces <- function(space, query_args, dest_folder) {
dir.create(dest_folder, recursive = TRUE, showWarnings = FALSE)
files <- tflow_files(space, query_args = query_args)
if (length(files) > 0) {
dest_paths <- file.path(dest_folder, basename(files))
file.copy(files, dest_paths, overwrite = TRUE)
message("Copied ", length(files), " files to ", dest_folder)
}
invisible(dest_paths)
}
# Download all fsLR 32k surfaces
download_surfaces("fsLR", list(density = "32k"), "~/my_surfaces/fsLR_32k")Common workflows
-
Vertex-wise stats: build your own
NeuroSurfacefromload_surface_template()and write stats intodata. -
Parcel summaries: use volumetric data with
reduce_atlas()or surface atlases withmap_atlas()to project parcel statistics back onto the mesh for visualization.
Dense vertex-wise overlays
plot_brain() supports dense (per-vertex) overlays in
addition to parcel-level colouring. You can pass a list with
lh and rh numeric vectors — one value per
vertex — and the result is a continuous colour map draped over the
cortical surface.
atl <- schaefer_surf(200, 7, space = "fsaverage6", surf = "inflated")
make_spatial_overlay <- function(atlas_hemi) {
verts <- neurosurf::vertices(neurosurf::geometry(atlas_hemi))
y <- verts[, 2]
z <- verts[, 3]
vals <- sin(y / 25) * cos(z / 30)
as.numeric(vals)
}
ov <- list(
lh = make_spatial_overlay(atl$lh_atlas),
rh = make_spatial_overlay(atl$rh_atlas)
)
plot_brain(
atl,
overlay = ov,
overlay_alpha = 0.7,
overlay_palette = "vik",
interactive = FALSE
)
Automatic NeuroVol projection
When overlay is a NeuroVol (e.g. a
whole-brain statistical map), plot_brain() automatically
projects it onto the surface via neurosurf::vol_to_surf() —
no manual preprocessing required. Here we build a synthetic volume with
two positive “activation” blobs over frontal cortex and one negative
blob over occipital cortex.
spacing <- c(4, 4, 4)
origin <- c(-80, -120, -60)
dims <- c(40L, 50L, 40L)
sp <- neuroim2::NeuroSpace(dims, spacing = spacing, origin = origin)
arr <- array(0, dim = dims)
for (i in seq_len(dims[1])) {
for (j in seq_len(dims[2])) {
for (k in seq_len(dims[3])) {
coord <- origin + (c(i, j, k) - 1) * spacing
d1 <- sum((coord - c(-20, 30, 40))^2)
d2 <- sum((coord - c( 20, 30, 40))^2)
d3 <- sum((coord - c( 0,-60, 20))^2)
arr[i, j, k] <- 3 * exp(-d1 / 800) +
3 * exp(-d2 / 800) -
2.5 * exp(-d3 / 600)
}
}
}
stat_vol <- neuroim2::NeuroVol(arr, sp)
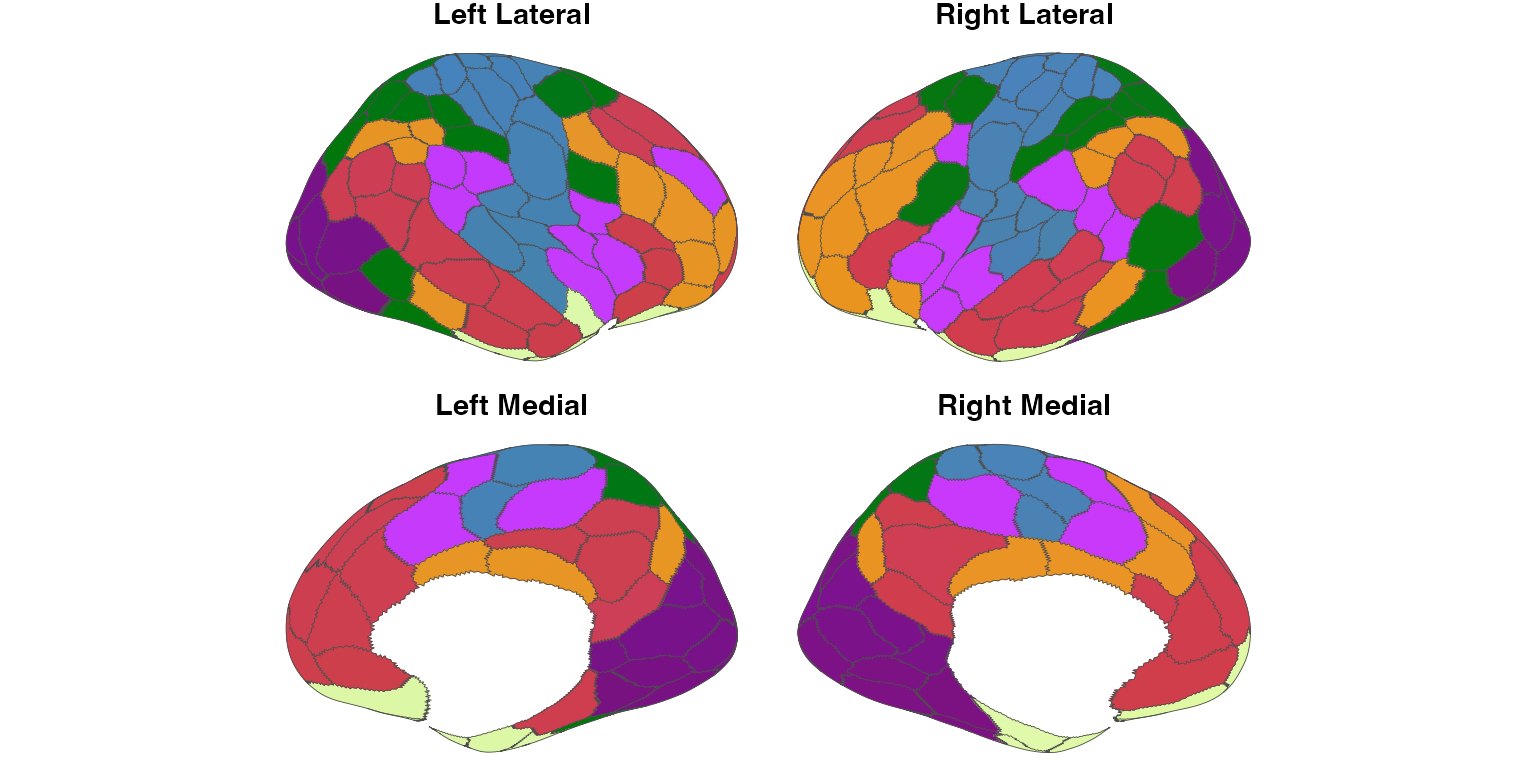
plot_brain(
atl,
overlay = stat_vol,
overlay_threshold = 0.5,
overlay_palette = "vik",
overlay_alpha = 0.7,
interactive = FALSE
)
In practice you would load a real statistical map instead:
stat_vol <- neuroim2::read_vol("my_tstat.nii.gz")
plot_brain(atl, overlay = stat_vol, overlay_threshold = 2,
overlay_palette = "vik", interactive = FALSE)Two optional parameters control the volume-to-surface projection:
-
overlay_fun: interpolation function —"avg"(default),"nn"(nearest-neighbour), or"mode"(most frequent label). -
overlay_sampling: sampling strategy —"midpoint"(default, halfway between white and pial),"normal_line"(multiple samples along the surface normal), or"thickness"(cortical-thickness–aware sampling).
plot_brain(
atl,
overlay = stat_vol,
overlay_fun = "nn",
overlay_sampling = "normal_line",
overlay_threshold = 2,
interactive = FALSE
)