Multi-Domain Graph Alignment with cone_align_multiple
Source:vignettes/cone-align-multiple.Rmd
cone-align-multiple.RmdOverview
cone_align_multiple() extends CONE-Align to three or
more graph domains by iteratively aligning each domain to a chosen
reference graph. The algorithm repeatedly solves pairwise
Procrustes/assignment problems until node correspondences stabilise.
This vignette demonstrates the workflow on the shared
alignment_benchmark dataset.
Loading the Benchmark Domains
alignment_benchmark <- manifoldalign::alignment_benchmark
multi_domains <- lapply(alignment_benchmark$domains, function(dom) {
multidesign::multidesign(dom$x, dom$design)
})
multi_names <- names(multi_domains)
node_counts <- vapply(multi_domains, function(dom) nrow(dom$x), integer(1))
hd_multi <- multidesign::hyperdesign(multi_domains)Running cone_align_multiple
cone_multi <- cone_align_multiple(
hd_multi,
ref_idx = 1,
ncomp = 6,
sigma = 0.9,
lambda = 0.05,
solver = "linear",
max_iter = 40,
tol = 1e-3
)
str(cone_multi, max.level = 1)
#> List of 10
#> $ v : num [1:12, 1:6] 6.16 8.33 8.12 6.02 -5.89 ...
#> $ s : num [1:240, 1:6] 0.136 0.1405 0.0687 0.1189 0.1419 ...
#> $ sdev : num [1:6] 0.1068 0.1008 0.0893 0.0907 0.0883 ...
#> $ preproc : NULL
#> $ block_indices:List of 3
#> $ assignment :List of 3
#> $ rotation :List of 3
#> $ n_domains : int 3
#> $ ref_idx : num 1
#> $ iterations : int 3
#> - attr(*, "class")= chr [1:2] "cone_align_multiple" "multiblock_biprojector"The reference domain (domain 1) is aligned to itself with the identity permutation. For the remaining domains we evaluate how often aligned pairs share the same latent class—again, labels were not used during optimisation and serve purely as a diagnostic.
assignments <- cone_multi$assignment
ref_labels <- multi_domains[[1]]$design$condition
accuracy_values <- vapply(seq_along(multi_names)[-1], function(idx) {
mean(ref_labels[assignments[[idx]]] == multi_domains[[idx]]$design$condition)
}, numeric(1))
accuracy_tbl <- tibble(
domain = multi_names[-1],
condition_accuracy = accuracy_values
)
accuracy_tbl
#> # A tibble: 2 × 2
#> domain condition_accuracy
#> <chr> <dbl>
#> 1 domain2 0.875
#> 2 domain3 0.85Visualising All Domains
# As with the pairwise vignette, class labels are only used for evaluation and
# plotting; `cone_align_multiple()` itself is unsupervised.
score_tbl <- as_tibble(as.matrix(cone_multi$s), .name_repair = "minimal")
colnames(score_tbl) <- paste0("comp", seq_len(ncol(score_tbl)))
scores <- score_tbl %>%
mutate(
domain = rep(multi_names, times = node_counts),
condition = rep(ref_labels, length(multi_names))
)
ggplot(scores, aes(x = comp1, y = comp2, colour = condition, shape = domain)) +
geom_point(size = 2, alpha = 0.8) +
labs(title = "cone_align_multiple embeddings (first two components)",
x = "Component 1", y = "Component 2") +
theme_minimal()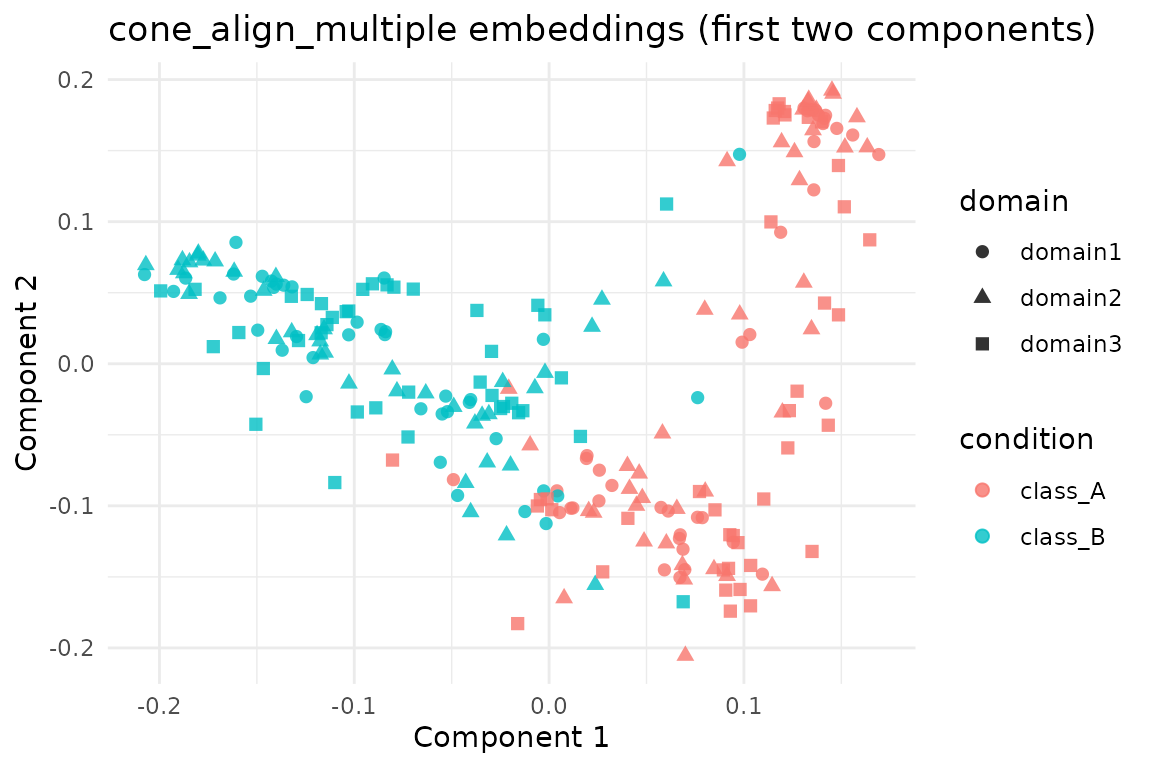
Summary
-
cone_align_multiple()aligns every domain to a shared reference using a sequence of pairwise CONE-Align steps. - On this benchmark the recovered correspondences preserve class labels across all domains (see the accuracy table above).
- Because the same dataset powers the
kemaandgpca_alignvignettes, you can now benchmark graph-vs-kernel-vs-linear alignment on identical inputs while recognising that the CONE variants remain unsupervised.