Advanced Regional MVPA Analysis in rMVPA
Bradley Buchsbaum
2026-05-08
Source:vignettes/Regional_Analysis.Rmd
Regional_Analysis.RmdIntroduction
Use regional MVPA when you already have an ROI mask and want one
prediction summary per region. The input is a 4D neuroimaging dataset
plus a design table; the output is a regional_mvpa_result
containing cross-validated performance, trial-level predictions, and
map-ready regional summaries.
This vignette follows one compact workflow: make a small dataset, define three ROIs, fit a blocked cross-validated classifier, and inspect the region-level performance.
Data Generation and Preparation
We begin by generating a synthetic volumetric dataset using the
gen_sample_dataset() function. This function creates a 4D
array (with spatial dimensions and multiple observations), along with a
binary mask and an associated design for cross-validation.
data_out <- rMVPA::gen_sample_dataset(D = c(6,6,6), nobs = 80, blocks = 4, nlevels = 2)
data.frame(
observations = length(data_out$design$block_var),
active_voxels = sum(data_out$dataset$mask),
blocks = length(unique(data_out$design$block_var)),
classes = length(levels(data_out$design$targets))
)
#> observations active_voxels blocks classes
#> 1 80 216 4 2The returned list contains two objects:
- dataset: an MVPA dataset object with training data and a binary mask.
- design: an MVPA design object specifying the response variable and block structure.
Setting Up the MVPA Model
Next, we create an MVPA model to evaluate a classification task. In
brief, we construct an mvpa_dataset, specify the design
(including the block variable and response), and define the model with
mvpa_model() using a chosen classifier and cross‑validation
strategy.
dset <- mvpa_dataset(data_out$dataset$train_data, mask = data_out$dataset$mask)
cval <- blocked_cross_validation(data_out$design$block_var)
mod <- load_model("sda")
tune_grid <- data.frame(lambda = 0.01, diagonal = FALSE)
mvpa_mod <- mvpa_model(
mod,
dataset = dset,
design = data_out$design,
crossval = cval,
tune_grid = tune_grid
)
mvpa_mod
#> mvpa_model object.
#> model: sda
#> model type: classification
#> tune_reps: 15
#> tune_grid:
#> lambda diagonal
#> 1 0.01 FALSE
#>
#> Blocked Cross-Validation
#>
#> - Dataset Information
#> - Observations: 80
#> - Number of Folds: 4
#> - Block Information
#> - Total Blocks: 4
#> - Mean Block Size: 20 (SD: 0 )
#> - Block Sizes: 1: 20, 2: 20, 3: 20, 4: 20
#>
#>
#> MVPA Dataset
#>
#> - Training Data
#> - Dimensions: 6 x 6 x 6 x 80 observations
#> - Type: DenseNeuroVec
#> - Test Data
#> - None
#> - Mask Information
#> - Areas: TRUE : 216
#> - Active voxels/vertices: 216
#>
#>
#> MVPA Design
#>
#> - Training Data
#> - Observations: 80
#> - Response Type: Factor
#> - Levels: a, b
#> - Class Distribution: a: 40, b: 40
#> - Test Data
#> - None
#> - Structure
#> - Blocking: Present
#> - Number of Blocks: 4
#> - Mean Block Size: 20 (SD: 0 )
#> - Split Groups: NoneThe model object packages the dataset, design, classifier, tuning grid, and cross-validation plan.
Running the Regional Analysis
The regional analysis is executed with run_regional().
Internally it prepares ROI indices from the region mask, applies the
MVPA model to each ROI, and then compiles performance metrics and
prediction tables.
regional_results <- run_regional(mvpa_mod, region_mask)The output is a regional_mvpa_result object with a
performance_table (cross‑validated metrics per region), a
prediction_table (trial‑level predictions), and
vol_results (volumetric maps of performance across the
brain).
Examining the Results
We can inspect the performance table to evaluate model accuracy in each region.
regional_results$performance_table
#> # A tibble: 3 × 3
#> roinum Accuracy AUC
#> <int> <dbl> <dbl>
#> 1 1 0.575 0.188
#> 2 2 0.488 0.00125
#> 3 3 0.488 -0.157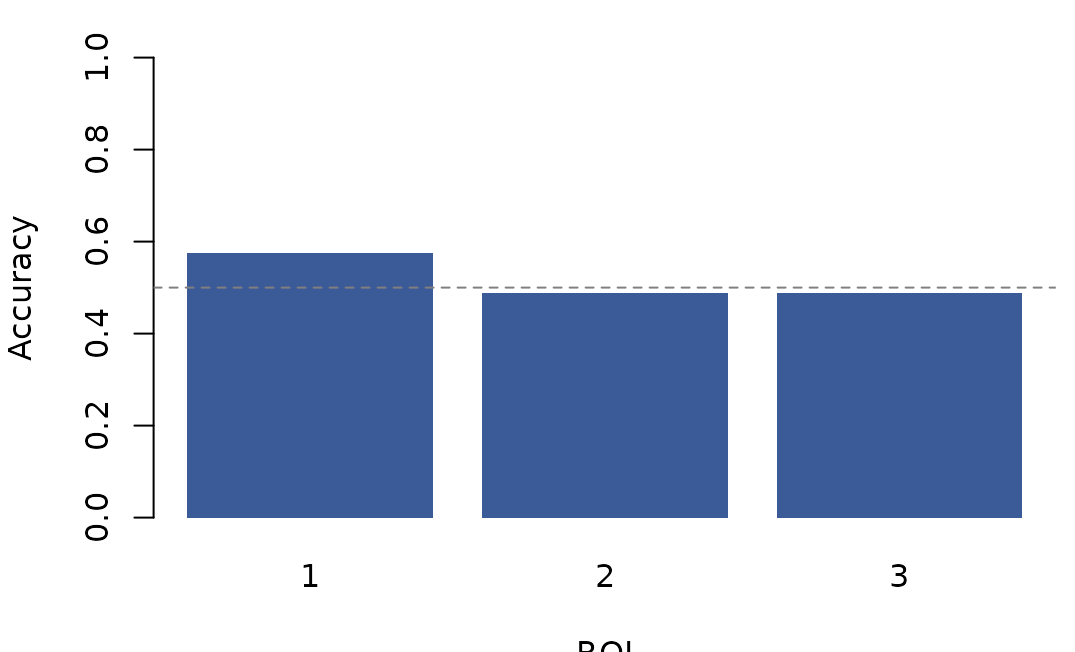
Cross-validated accuracy per region. The dashed line marks chance for a 2-class problem.
The bar plot makes between-region differences visible at a glance and shows where each region sits relative to chance. For a compact trial-level check, show the first few predictions:
utils::head(regional_results$prediction_table)
#> # A tibble: 6 × 8
#> # Rowwise:
#> .rownum roinum observed pobserved predicted correct prob_a prob_b
#> <int> <int> <fct> <dbl> <fct> <lgl> <dbl> <dbl>
#> 1 1 1 b 0.872 b TRUE 0.128 0.872
#> 2 2 1 a 0.961 a TRUE 0.961 0.0393
#> 3 3 1 a 0.996 a TRUE 0.996 0.00371
#> 4 4 1 b 0.983 b TRUE 0.0166 0.983
#> 5 5 1 b 0.0243 a FALSE 0.976 0.0243
#> 6 6 1 a 0.805 a TRUE 0.805 0.195Volumetric results (vol_results) can be further
visualized with neuroimaging tools to determine spatial patterns of
performance.
Summary
This vignette showed you how to generate synthetic neuroimaging data, define ROIs with region masks, set up MVPA models with cross-validation, run analyses across ROIs, and interpret the performance metrics. You now have the tools to conduct regional MVPA analyses on your own neuroimaging data.
For further details, see
vignette("Searchlight_Analysis") for the searchlight
counterpart and vignette("CrossValidation") for
cross-validation strategies.